Crossref Citations
This article has been cited by the following publications. This list is generated based on data provided by
Crossref.
Katsuno, Hiroyasu
Kimura, Yuki
Yamazaki, Tomoya
and
Takigawa, Ichigaku
2022.
Early Detection of Nucleation Events From Solution in LC-TEM by Machine Learning.
Frontiers in Chemistry,
Vol. 10,
Issue. ,
Kimura, Yuki
Katsuno, Hiroyasu
and
Yamazaki, Tomoya
2022.
Possible embryos and precursors of crystalline nuclei of calcium carbonate observed by liquid-cell transmission electron microscopy.
Faraday Discussions,
Vol. 235,
Issue. ,
p.
81.
Ide, Yuki
Shirakura, Hayato
Sano, Taichi
Murugavel, Muthuchamy
Inaba, Yuya
Hu, Sheng
Takigawa, Ichigaku
and
Inokuma, Yasuhide
2023.
Machine Learning-Based Analysis of Molar and Enantiomeric Ratios and Reaction Yields Using Images of Solid Mixtures.
Industrial & Engineering Chemistry Research,
Vol. 62,
Issue. 35,
p.
13790.
Li, WeiDong
Xie, Bo
Meng, Chunmei
and
Hu, Yanling
2023.
Improving the Quality of Unstained Transmission Electron Microscope Images Using Deep Learning.
p.
1.
Lyu, Zhiheng
Yao, Lehan
Chen, Wenxiang
Kalutantirige, Falon C.
and
Chen, Qian
2023.
Electron Microscopy Studies of Soft Nanomaterials.
Chemical Reviews,
Vol. 123,
Issue. 7,
p.
4051.
KIMURA, Yuki
KATSUNO, Hiroyasu
HIRAKAWA, Shizuka
and
YAMAZAKI, Tomoya
2023.
Data-driven “<i>In situ</i>” Liquid-cell Transmission Electron Microscope Observation by Machine Learning.
Vacuum and Surface Science,
Vol. 66,
Issue. 12,
p.
700.
Chen, K
and
Barnard, A S
2024.
Advancing electron microscopy using deep learning.
Journal of Physics: Materials,
Vol. 7,
Issue. 2,
p.
022001.
Katsuno, Hiroyasu
Kimura, Yuki
Yamazaki, Tomoya
and
Takigawa, Ichigaku
2024.
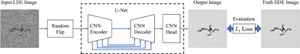
Machine Learning Refinement of In Situ Images Acquired by Low Electron Dose LC-TEM.
Microscopy and Microanalysis,
Vol. 30,
Issue. 1,
p.
77.
Hata, Satoshi
Ihara, Shiro
Saito, Hikaru
and
Murayama, Mitsuhiro
2024.
In-situ heating-and-electron tomography for materials research: from 3D (in-situ 2D) to 4D (in-situ 3D).
Microscopy,
Vol. 73,
Issue. 2,
p.
133.
Kim, Joodeok
Kang, Sungsu
Cheng, Fanrui
Wang, Yi
Ye, Xingchen
and
Park, Jungwon
2024.
Recent advances in liquid phase transmission electron microscopy of nanoparticle growth and self-assembly.
MRS Bulletin,
Vol. 49,
Issue. 4,
p.
365.
 $5$
$5$  $e^{-}$ per pixel becomes comparable to that of images acquired with a total dose of approximately
$e^{-}$ per pixel becomes comparable to that of images acquired with a total dose of approximately  $1{,}000$
$1{,}000$  $e^{-}$ per pixel. Because the conversion time is approximately 8 ms, in situ observation at 125 fps is possible. This imaging technique enables in situ observation of electron-beam-sensitive specimens.
$e^{-}$ per pixel. Because the conversion time is approximately 8 ms, in situ observation at 125 fps is possible. This imaging technique enables in situ observation of electron-beam-sensitive specimens.