Implications
No direct evidence has been reported about animal health effects of genetically modified (GM) feed. Although the commercialisation of several GM feeds, including RoundUp Ready®soybean, has been authorised by the European authorities, concerns over safety still persist in the public, mainly regarding either the detection of transgenic plant genes and proteins in animal systems or allergenicity and toxicity of GM plants. The aims of this study were to evaluate the presence of plant DNA fragments and the levels of some health parameters in animals fed on GM soybean.
Introduction
In recent years, genetic engineering has been widely applied to agriculture in order to obtain specific plant characteristics, which can lead to an improvement in both food quality and production. In this context, the European Food Safety Authority has stated that no recombinant DNA sequences have been found in any organ or tissue sample from animals fed genetically modified (GM) plants (EFSA, 2007). On this basis, a number of GM products have been approved for human consumption, but concerns over safety persist in the public, mainly regarding either the detection of transgenic plant genes and proteins in animal systems or allergenicity and toxicity of GM plants. Furthermore, some authors have shown the possible survival of recombinant plant DNA fragments after digestion, although health implications as well as biological relevance of data are still far from being confirmed (Nemeth et al., Reference Nemeth, Wurz, Artim, Charlton, Dana, Glenn, Hunst, Jennings, Shilito and Song2004; Mazza et al., Reference Mazza, Soave, Morlacchini, Piva and Marocco2005; Sharma et al., Reference Sharma, Damgaard, Alexander, Dugan, Aalhus, Stanford and McAllister2006). The absorption of plant DNA across the intestinal barrier is a natural event, as demonstrated by the detection of endogenous plant genes in several animal tissues and products (Reuter and Aulrich, Reference Reuter and Aulrich2003; Tony et al., Reference Tony, Butschke, Broll, Grohmann, Zagon, Halle, Dänicke, Schauzu, Hafez and Flachowsky2003; Tudisco et al., Reference Tudisco, Lombardi, Bovera, D’Angelo, Cutrignelli, Mastellone, Terzi, Avallone and Infascelli2006b). In a preliminary study (Laudadio et al., Reference Laudadio, Petrera, Tudisco and Infascelli2006) concerning possible plant DNA fragment survival after digestive processes and their transfer and accumulation in tissues and organs of offspring, we found the chloroplast sequence, but no transgenic fragments in blood and tissues from lambs fed dam’s milk until weaning. No relations between GM food and health have been reported to date.
Many feeding trials have been reported in which GM foods have been administered to rats or mice for prolonged periods, and parameters such as body weight (BW), feed consumption, blood chemistry, organ weights and histopathology have been measured. The majority of these experiments indicated no clinical effects nor histopathological abnormalities in organs or tissues of exposed animals (EFSA, 2008). By contrast, Malatesta et al. (Reference Malatesta, Caporaloni, Gavaudan, Rocchi, Serafini, Tiberi and Gazzanelli2002) found significant nuclear modifications of hepatocytes in young and adult (2 to 8 months of age) female mice fed on GM soybean. In particular, GM-fed mice show irregularly shaped nuclei, which generally represents an index of high metabolic rate, and a higher number of nuclear pores, suggesting intense molecular trafficking. More recently, the same authors (Malatesta et al., Reference Malatesta, Boraldi, Annovi, Baldelli, Battistelli, Biggiogera and Quaglino2008) showed that several proteins belonging to hepatocyte metabolism, stress response, calcium signalling and mitochondria were expressed in 24-month-old female mice fed on GM soybean, indicating a more marked expression of senescence markers in comparison to controls. Moreover, hepatocytes of GM-fed mice showed mitochondrial and nuclear modifications indicative of reduced metabolic rates. Furthermore, Tudisco et al. (Reference Tudisco, Infascelli, Cutrignelli, Bovera, Morcia, Faccioli and Terzi2006a) hypothesised that cell metabolism of several enzymes was altered in rabbit fed on GM soybean.
The aims of this study were to investigate the presence of plant DNA fragments in blood and milk from goats fed GM soybean and in tissues and organs from their kids fed only mother’s milk for 60 days, and to study the activity of organ-specific enzymes.
Material and methods
Diets, animals and feeding
Twenty pregnant dairy goats were equally assigned to control (C) and treated (T) groups, homogeneous in parity and milk production at the previous lactation. Animals from different groups were housed in separate sheds. Experiments started 2 months before kidding; the animals were fed a diet consisting of oat hay and concentrate [crude protein (CP) 18% of dry matter (DM); metabolisable energy (MJ/DM) 12.22], the latter containing solvent extracted (s.e.) soybean meal (13% of concentrate DM) which was from conventional or GM (RoundUp Ready®(RR)) soy beans for groups C and T, respectively. RR is tolerant to the glyphosate family of herbicides by expressing transgenic DNA from the CP4 strain of Agrobacterium tumefaciens (CP4 EPSPS), encoding 5-enolpyruvilshikimate-3-phosphate synthase protein (glyphosate-tolerant soybean GTS 40-3-2; Padgette et al., Reference Padgette, Kolacz, Delannay, Re, LaVallee, Tinius, Rhodes, Otero, Barry, Eichholtz, Peschke, Nida, Taylor and Kishore1995). A polymerase chain reaction (PCR)-end point reaction confirmed the presence of p35S and CP4 epsps specific transgenes in the treated diet as well as their absence in the control diet. Both groups received oat hay ad libitum, while the concentrate was administered in amounts of 200, 300 and 400 g/head per day, 60, 30 and 15 days before kidding, respectively. After kidding, administration of concentrate was gradually increased up to 700 g/head per day and oat hay was replaced by alfalfa hay in order to increase diet protein content. Water was given ad libitum. The chemical composition (Van Soest et al., Reference Van Soest, Robertson and Lewis1991; AOAC, 2000) and metabolisable energy (INRA, 1978) of hays and concentrate as well as ingredients of concentrate are reported in Table 1.
Table 1 Chemical composition (g/kg DM) and ME (MJ/kg DM) of hays and concentrate
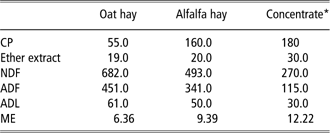
DM = dry matter; ME = metabolisable energy.
*Ingredients (g/Kg DM): soft wheat bran 30; soybean s.e. 13; corn meal 13; sunflower meal 10.5; citrus pulp 8; sugar beet pulp 7.9; corn gluten feed 7; sugarcane molasses 7.5; CaCO3 1.5; CaHPO4 0.7; vitamins 0.2; NACL 0.7.
All goats had twin deliveries each of them with at least one male kid. Immediately after kidding, 10 male kids, were randomly selected from each group and allowed into individual cages positioned in a separate room. Each kid was fed only milk from its mother using a milk feeder. Milk was collected from each goat into sterile tubes with aseptic techniques and administered twice/day until 12 h before slaughtering. Kids were slaughtered at 60 ± 7 days of age (11.2 ± 1.1 kg live weight). Kid BWs were taken at birth and immediately before slaughtering.
Sample collection
In all, 20 dams (ten per group) with their kids were included in the trial. After kidding, 100 ml of milk from each goat (obtained by mixing the production of the two daily milkings) were collected at days 15, 30, 45, 60 and 75. Samples were immediately frozen on dry ice before storage at −20°C. Blood was withdrawn every month (day −60 until day 60) from each goat via jugular vein puncture. Blood was removed from kids before slaughter. In both cases, samples used for DNA extraction were added to tubes containing K3-EDTA (ethylene diamine tetra acetic acid) 7.5%. Before inserting the needle, the surface was cleaned and the first millilitre of blood was discarded in order to avoid contamination.
Organs and tissues from kids (liver, spleen, kidney, heart and skeletal muscle) were removed. Slaughter and sampling rooms were close, but distinct in order to avoid possible contamination. For the same reason, the outermost layer of tissue from each organ was removed to sample the inner part. Three aliquots (25 mg each) from each sample were immediately stored at −20°C in sterile tubes for DNA extraction. Later on, each aliquot underwent PCR reaction. Specimens from kidney, liver, heart and muscle from right leg samples (5 g each) were washed in saline and stored at −80°C to determine enzyme activity.
DNA extraction and quantification
Milk (10 ml) and, as control, conventional and transgenic soybean meal s.e. samples (100 mg) were extracted in duplicate according to the Wizard extraction method (Promega, Madison, WI, USA). Milk samples were incubated at 4°C overnight and centrifuged (2.000×g for 20 min at 4°C) to separate cream, skimmed milk and sediment; only the sediment fraction was subjected to DNA extraction. The somatic cell pellet obtained was washed twice with phosphate buffered saline (PBS, pH 7.2). The pellet and the ground plant samples were resuspended by vortexing in 860 μl of extraction buffer [10 mM Tris-HCl (pH 8.0), 150 mM NaCl, 2 mM EDTA, 1% (w/v) SDS], 100 μl guanidine hydrochloride (5M) and 40 μl of proteinase K (20 mg/ml) and then incubated at 58°C for at least 3 h on a shaking incubator and centrifuged at 20.000×g for 10 min. Five hundred microlitres of the supernatant were incubated with 5 μl RNAse (10 mg/ml) at 37°C for 10 min. One millilitre of Wizard DNA purification resin (Promega) was added to the supernatant. The DNA-resin mixture was pushed through the column and washed with 2 ml 80% (v/v) isopropyl alcohol followed by centrifugation at 20.000×g for 1 min. After drying at 70°C for 10 min, the DNA was eluted with 50 μl of 70°C elution buffer [10 mM Tris-HCl (pH 9.0), 0.1 mM EDTA] and centrifuged at 20.000×g for 1 min.
NucleoSpin® Tissue and NucleoSpin® Blood kit (Macherey-Nagel, Duren, Germany) were used for extraction of tissue (25 mg) and blood (200 μl) samples, respectively, according to the manufacturer’s protocol. Each sample was extracted in duplicate and stored at −20°C until use. In such a way, a total of six samples for each kid tissue and organ were obtained. In addition, as suggested by Nemeth (2004), negative (buffer only) control to each set of DNA extraction was included to monitor for any contamination that could have occurred during the DNA extraction procedure. The DNA was quantified by spectrophotometer (Thermo Electron Corporation, Waltham, MA, USA) according to standard molecular techniques (Sambrook et al., Reference Sambrook, Fritsh and Maniatis1989).
PCR analyses
In order to avoid contamination, PCR reactions were assembled in an ultraviolet-sterilised hood. Filter tips against sample aerosol and sterile disposable tubes were used during pipetting. All PCR amplifications were performed on a Gene Amp PCR System 2400 (Applied Biosystems, Foster City, CA, USA). DNA was amplified using Taq DNA polymerase (Invitrogen, Carlsbad, CA, USA) in accordance with the manufacturer’s instructions. Sequence, amplicon size and annealing temperature of the primer pair sets (Sigma-Genosys Ltd, Haverhill, UK) used for PCR are shown in Table 2.
Table 2 Sequence (5′-3′), amplicon size (bp) and annealing temperature (°C) of primer pairs used in PCR

Primers Cap 144/496 were previously used to amplify a conserved portion of caprine mtDNA, which encodes the 12S ribosomal RNA (12S rRNA) gene of mitochondrial DNA from caprine (Bottero et al., Reference Bottero, Civera, Numera, Rosati, Sacchi and Turi2002).
Subsequently, samples were monitored for the presence of the chloroplast sequence for tRNA Leu by using the Clor 1/2 primers designed on the chloroplast trnL sequence (Terzi et al., Reference Terzi, Infascelli, Tudisco, Russo, Stanca and Faccioli2004). The obtained amplicon was analysed by electrophoretic separation and purified from gel. The identity was verified by sequencing and subsequently by Basic Local Alignment Search Tool (BLAST) analysis.
About the sensibility, we performed a real time PCR with Clor1/Clor2 primer and specific TaqMan probe to verify the presence of tRNA Leu gene, housekeeping gene in conventional and transgenic genotype. Then, we elaborated a standard curve and the results obtained demonstrated the efficiency of primer/probe combination to notice the presence of soybean genotype and to give quantification.
Finally, species-specific primers for conventional and GM soybean were used: Le1n02 5/3 which amplifies the soybean lectin gene (Kuribara et al., Reference Kuribara, Shindo, Matsuoga, Futo, Aoki, Hirao, Ariyama, Goda, Toyoda and Hino2002); 35S 1/2 and CP4 EPSPS 1/2 which amplify, respectively, part of the 35S promoter (Lipp et al., Reference Lipp, Brodmann, Pietsch, Pauwels and Anklam1999) derived from the cauliflower mosaic virus and part of the specific gene sequence (CP4 epsps) that provides herbicide tolerance derived from Agrobacterium tumefaciens strain CP4 both present in the genomic structure of GM soybean (Hernández et al., Reference Hernández, Rodríguez-Lázaro, Esteve, Prat and Pla2003). The primer pairs were selected from those reported in the literature (Jennings et al., Reference Jennings, Kolwyck, Kats, Whetsell, Surber, Cromwell, Lirette and Glenn2003) with the aim of obtaining short amplicons (118 bp), compatible with highly fragmented DNA samples.
A 100 base pairs (bp) ladder, containing linear DNA fragments, served as size standard reference. The PCR was done three times, and samples with positive results at least twice were judged as positive (Chowdhury et al., Reference Chowdhury, Mikami, Nakajima, Kuribara, Hino, Suga, Hanazumi and Yomemochi2003b). In every PCR run, positive and negative controls were included to ensure reproducibility and absence of contaminants. For positive controls, reference DNA consisting of purified conventional and transgenic soybean DNA was amplified in parallel with the samples to ensure correct performance of the PCR; for negative control (buffer blank), water instead of DNA was added to the PCR mix to check for cross contamination with soybean DNA in the PCR mix or its constituents (Klaften et al., Reference Klaften, Whetsell, Webser, Grewal, Fedyk, Einspanier, Jennings, Lirette and Glenn2004).
Enzyme assay
The levels of aspartate aminotransferase (AST), alanine aminotransferase (ALT), creatine kinase, lactic dehydrogenase (LDH), gamma glutamyltransferase (GGT) and alkaline phosphatase were determined in serum and homogenates prepared from samples of liver, kidney, heart and skeletal muscle. Briefly, tissue (1 g) was homogenised in ice-cold homogenisation buffer (in mM): 280 mannitol, 10 KCl, 1 MgCl2, 0.2 Pefabloc SC, 10 Hepes, pH 7.0 adjusted to pH 7 with Tris HCl 10 mM. After centrifugation at 10 000×g for 10 min the upper layer was used for analysis.
Enzyme activity was determined spectrophotometrically (340 nm) by using reagents from Spinreact SA (Sant Esteve de Bas, Spain). Determinations were performed according to the manufacturer’s instructions, including the convertion of the results into units/litre. In order to assess the isoenzymatic distribution of LDH, electrophoretic separation was performed on each sample. Briefly, 20 μl of sample were applied on cellulose acetate membranes and electrophoresis was performed under denaturing conditions at 200 V for 50 min in barbital buffer. The substrate (tris pH 7.4, sodium lactate, sodium azide) was incorporated in agarose and put on the acetate membrane immediately after electrophoresis. Therefore, it was possible to observe LDH isoenzymes (ISO-LAD commercial kits by Chemetron Chimica S.p.A., Milan, Italy). Percentage of isoenzyme fractions of total LDH were quantified by using a densitometer (CGA, Florence, Italy). Data were then converted into units/litre (serum) or units/gram of tissue (organs).
LDH histochemistry
Samples were prepared using a cryostat: 10 μm sections were obtained, with five serial sections put in every slide. The frozen sections were air-dried and then incubated, always in the dark, in LDH-incubating medium at 37°C for 10 min. The incubated medium consisted of 50 mM Tris-HCL (pH 7.4) containing 67 mM sodium lactate (L 1375, Sigma-Aldrich, Milan, Italy), 5 mM nitroblue tetrazolium (N 6876, Sigma), 5 mM MgCl2 and 3 mM NAD (N1511, Sigma). Each sample, both from control and treated animals, was compared with its negative control section, which was obtained by incubation in incubating medium without sodium lactate. The histochemical reaction was stopped, at the same time for every slide, by fixing the section in formalin for 15 min at room temperature. The sections were then counterstained with methyl green. The slides were mounted in PBS/glycerol (1 : 1) and then examined and photographed. Five slides for each examined tissue were independently evaluated by two observers using a DMRA2 microscope (Leica Microsystems Wetzlar GmbH, Germany).
Statistics
Differences in kid slaughter weight as well as spleen, liver, kidney and heart were tested by one way analysis of covariance (control v. treated group). Data were covariated for birth weight to reduce variations from the mean square error in the dependent variables. For goat milk and blood, differences in gene detection among sampling periods were tested by the Cochran-Q test. The presence of plant DNA fragments in kids blood, skeletal muscle and organs was analysed by using the χ 2 test. Results for enzyme assays were expressed as mean ± standard deviation and differences between groups were analysed by the Student’s t-test. All statistics were performed with SPSS 12.1 software (SPSS, 1999).
Results
BW of the kids at birth was on average 4.2 v. 3.7 kg, respectively for groups C and T. As a consequence, birth weight was used to covariate slaughter and organ weights. Neither body nor organ weights significantly differed between groups (Table 3).
Table 3 Body and organ weights in control (n = 10) and treated (n = 10) kids fed RoundUp Ready®soybean meal

All data were collected at slaughter.
Detection of DNA fragments
The quality of each DNA sample extracted from milk and blood of control and treated goats and from liver, kidney, skeletal muscle, spleen, heart and blood of each group of kids was first verified using the Cap 144/496 primers that were used to amplify a conserved portion of caprine mtDNA 12S rRNA sequence. An example is reported in Figure 1.

Figure 1 Representative electrophoresis gels of amplified DNA in: (left) milk and blood from control (lines 1 and 3) and treated (lines 2 and 4) goats; (right) liver, kidney, skeletal muscle, spleen, heart and blood from control (lines 1 to 6, respectively) and treated (lines 7 to 12, respectively) kids. In each panel, lane M contains a 100 bp DNA ladder; ‘−’ is a negative control (no DNA template).
Table 4 shows the number of goats in which different DNA sequences were detected. PCR was done three times for each sample and only samples with at least twice a positive result were judged as positive (Chowdhury et al., Reference Chowdhury, Mikami, Nakajima, Kuribara, Hino, Suga, Hanazumi and Yomemochi2003b). A chloroplast specific DNA fragment (trnL, 100 bp) was found, both in blood and milk, in the majority of control and treated samples. In the samples, which were positive for the presence of chloroplast DNA, the fragments of the lectin gene, 35S promoter and CP4 epsps were investigated and, again, several control and treated samples were found positive for lectin gene fragments. Transgenic target DNA sequences (35S and CP4 epsps) were not detected in blood and milk from control goats that received a diet containing conventional soybean meal. By contrast, transgenic DNA fragments were amplified from samples (blood and milk) from goats that received transgenic soybean. The Cochran’s Q-test revealed that detection of chloroplast, lectin, 35S promoter and CP4 epsps gene fragments in goats’ milk and blood was significant (P < 0.01) during the whole experiment.
Table 4 Number of animals in which DNA sequences were detected. Blood and milk derived from dams which either (treated, n = 10) or not (control, n = 10) were fed Roundup Ready soybean meal s.e.

chlor = chloroplast DNA fragment.
**P < 0.01.
Table 5 shows the number of kids from control and treated group in which different DNA sequences were detected. The chloroplast specific DNA fragment (trnL, 100 bp) was found, in all organs, in the majority of control and treated samples. In the samples, which proved positive for the presence of chloroplast DNA, the fragments of lectin gene, 35S promoter and CP4 epsps were investigated and, again, several control and treated samples were found positive for lectin gene fragments. Transgenic target DNA sequences (35S and CP4 epsps) were not detected in organs from control kids whose mothers received a diet containing conventional soybean meal. In contrast, transgenic DNA fragments were amplified from some samples of kids whose mothers received transgenic soybean. As seen, the CP4 epsps fragment was found in more samples compared with the 35S fragment, the reason for such result is not clear, but it could be due to slight differences in PCR conditions. In order to assess the significance of the presence of transgenic DNA fragments in kids organs, a χ 2 analysis of all kids data v. those detected in milk from the corresponding mothers at day 60 (slaughtering time) was performed and revealed that: (i) chloroplastid fragment presence was significant in all organ and tissues from both control and treated groups, (ii) lectin fragments detection was significant in kidney and liver of both groups (P < 0.01), in blood and muscle from control group (P < 0.05) and in blood, muscle and heart from treated group (P < 0.05), (iii) the detection of 35S promoter was significant (P < 0.01) in liver, kidney and blood of treated kids and (iv) the detection of CP4 epsps gene fragment was significant in liver and kidney (P < 0.05) and in heart and muscle (P < 0.01) from treated group.
Table 5 Number of dams producing milk at 60 days in lactation in which DNA sequences were detected. Number of kids with organs in which DNA sequences were detected

Milk derived from dams which either (treated, n = 10) or not (control, n = 10) were fed Roundup Ready soybean meal s.e. Organs derived from their nursing kids. The lectine, 35S and CP4 epsps fragments were investigated only in those samples, which were positive for the chloroplast DNA fragment (Chlor).
*P < 0.05; **P < 0.01. This P-value indicates a significant proportion of the kids found positive for the same DNA fragments detected in milk.
In Figure 2, a representative example is presented illustrating the detection of endogenous high copy chloroplast specific DNA fragment (trnL, 100 bp) in goats milk and blood of both groups and in tissue, organs and blood from both control and treated groups of kids.

Figure 2 Confirmation of amplified endogenous chloroplast specific DNA fragments, trnL (representative data) in: (left) milk and blood from control (lines 1 and 3) and treated (lines 2 and 4) goats; (right) liver, kidney, skeletal muscle, spleen, heart and blood from control (lines 1 to 6, respectively) and treated (lines 7 to 12, respectively) kids. In each panel, lane M contains a 100 bp DNA ladder; ‘−’ is a negative control (no DNA template) and ‘+’ is a positive control (DNA extracted from soybean meal).
In Figure 3, a representative example obtained by using Le1n02 5/3 soybean specific primers shows the lectin signal detected in goats blood and milk samples and in the plant control, as well as in control and treated groups of kids.

Figure 3 Representative electrophoresis analysis of single copy specific soybean DNA fragment, lectin gene in: (left) milk and blood from control (lines 1 and 3) and treated (lines 2 and 4) goats; (right) liver, kidney, skeletal muscle, spleen, heart and blood from control (lines 1 to 6, respectively) and treated (lines 7 to 12, respectively) kids. In each panel, lane M contains a 100 bp DNA ladder; ‘−’ is a negative control (no DNA template), and ‘+’ is a positive control (DNA extracted from soybean meal).
Transgenic target DNA sequences (35S and CP4 epsps) detected in either goats that received a diet containing transgenic soybean meal or their kids as well as in the RoundUp Ready soybean positive control are represented in Figure 4. In particular, in blood and milk of treated goats, Figure 4 shows fragments of 35S promoter and fragments of the CP4 epsps gene. In treated kids, a representative detection of fragments of 35S and the CP4 epsps gene is reported.

Figure 4 Representative data of amplified transgenic DNA fragments. (a) 35S promoter fragments (195 bp) in milk and blood from control (lines 1 and 2, respectively) and treated (lines 3 and 4, respectively) goats. (b) 35S promoter fragments (195 bp) in liver, kidney, spleen, heart, skeletal muscle and blood from control (lines 7 to 12, respectively) and treated (lines 1 to 6, respectively) kids. (c) fragments of CP4 epsps gene (145 bp) in milk and blood from control (lines 3 and 4, respectively) and treated (lines 1 and 2, respectively) goats. (d) fragments of CP4 epsps gene (145 bp) in liver, kidney, spleen, heart, skeletal muscle and blood from control (lines 1 to 6, respectively) and treated (lines 7 to 12, respectively) kids. In each panel, lane M contains a 100-bp DNA ladder, ‘−’ is a negative control (no DNA template), and ‘+’ is a positive control (DNA extracted from Roundup Ready soybean meal).
Since it could be likely that the DNA detected may be too small to be of any biological significance, we later investigated whether larger DNA fragments could be detected in milk samples. To this purpose, a primer amplifying a fragment of 447 bp of the p-35S/CP4 epsps gene (Cardarelli et al., Reference Cardarelli, Branquinho, Ferreira, da Cruz and Gemal2005) was used. In 57% of the samples that were found positive for smaller transgenic fragment, such sequence was detected (see addendum).
Enzyme activity
As depicted in Table 6, AST and ALT were significantly lower in serum from goats fed GM soybean at day 60 and at days 30 and 60, respectively. Overall, very limited differences in the monitored enzyme activity have been observed.
Table 6 Monthly levels of AST, ALT, CK, LDH, GGT and ALP in serum taken 60 days before to 60 days after lambing from control (C) and treated (T) group of goats

AST = aspartate aminotransferase; ALT = alanine aminotransferase; CK = creatine kinase; LDH = lactic dehydrogenase; GGT = gamma glutamyltransferase; ALP = alkaline phosphatase.
*P < 0.05 control (C) v. treated (T) dams.
Figure 5 shows the differences in enzyme activity in kids’ heart, skeletal muscle, kidney, liver and serum. Statistical differences (P < 0.05) were detected in kidney for GGT and LDH, whereas in the heart and skeletal muscle this result was seen only for LDH (P < 0.05). Hence, in five of the 30 cases presented, differences were observed.

Figure 5 Levels of creatine kinase (CK), lactic dehydrogenase (LDH), aspartate aminotransferase (AST), alanine aminotransferase (ALT), gamma glutamyltransferase (GGT) and alkaline phosphatase (ALP) in serum and in homogenates from liver, kidney, heart and skeletal muscle from control (![]() ) and treated (
) and treated (![]() ) kids. *P < 0.05.
) kids. *P < 0.05.
No statistical differences were found for studied enzymes in serum and liver. The increase in LDH activity was confirmed by histochemistry (Figure 6). LDH histochemistry showed a widespread distribution of enzyme activity in all the tissues that were examined. The expression of LDH activity was further illustrated by a violet staining in the cytoplasm of the cells. In tissue sections used as negative controls this staining was light blue. Myocytes, myocardiocytes, epithelial cells of renal tubules and hepatocytes all expressed LDH activity. LDH staining was more intense in myocytes, myocardiocytes and epithelial cells of renal tubules of treated than in those of control animals. Arrows show some zones where LDH histochemical staining is more intense. No difference was observed between the two groups of animals in hepatocytes.

Figure 6 LDH histochemistry on negative control sections (a, d, g, l), control (b, e, h, m) and treated (c, f, i, n) animals. LDH activity was revealed by a violet staining. In negative control sections, a light bluish staining was observed. In the skeletal muscle, heart and epithelial cells of the renal tubule of treated animal (c, f, i) the intensity of LDH staining (arrows) was higher than in the same tissues (b, e, h) of control animals. Bar = 50 μm.
Table 7 shows the relative distribution of LDH isoenzymes in serum and in tissues. Significant differences (P < 0.05) between control and treated animals were detected for heart and kidney LDH1 and for muscles LDH1 and LDH2, thus confirming the significant increase of the enzyme in these tissues.
Table 7 Relative distribution of LDH isoenzymes in serum (S) and in homogenates from heart (H), SM, kidney (K) and liver (L) from control (C) and treated (T) group of kids

LDH = lactic dehydrogenase; SM = skeletal muscle.
*P < 0.05 control (C) v. treated (T) dams.
In addition, analysis of isoenzyme distribution revealed that LDH1 significantly increased also in the liver if we compare the proportional distribution of isoenzymes (51.2 ±3.7 v. 57.8 ± 1.4). Moreover, heart LDH2 (4.4 ± 1.9 v. 2.9 ±0.9) and LDH3 (1.9 ± 0.4 v. 1.1 ± 0.7) significantly decreased and a similar result was seen for muscle LDH5 (55.6 ± 5.7 v. 49.4 ± 2.8).
Discussion
In our study, the weight of the organs of kids did not differ between groups, in accordance to findings of previous research (Tudisco et al., Reference Tudisco, Lombardi, Bovera, D’Angelo, Cutrignelli, Mastellone, Terzi, Avallone and Infascelli2006b) carried out on rabbits fed GM soybean meal.
Detection of DNA fragments
As stated by the EFSA (2008), several aspects have to be investigated when considering whether or not recombinant DNA from GM plants, or the derived proteins can end up in animal tissues and products. These include (i) the fate of the recombinant DNA and protein during feed processing and ensiling; (ii) the fate of the recombinant DNA and protein in the gastrointestinal tract of animals fed with GM feed; (iii) the potential absorption of the digested pieces of DNA or protein into animal tissues/products and (iv) the potential of biological functionality of absorbed DNA and protein fragments.
Our findings show plant DNA fragments are likely to survive digestive processes to some extent (Duggan et al., Reference Duggan, Chambers, Heritage and Forbes2003; Einspanier et al., Reference Einspanier, Lutz, Rief, Berezina, Zverlov, Schwarz and Mayer2004), as well as their transfer to blood and milk. In addition, the detection of plant DNA in tissues and organs of nursed kids could support the hypothesis of a gene transfer through milk. Chloroplast target DNA sequences were found in our previous research in blood and tissues of lambs allowed to nurse from the dam. However, no single copy feed fragment was detectable in lamb samples nor in the mother’s milk and blood. The reason for this high recovery of chloroplast DNA in animal tissues is a consequence of the larger quantities of these genes as compared with transgenic genes, which are usually found as a single copy gene (Aumaitre et al., Reference Aumaitre, Aulrich, Chesson, Flachowsky and Piva2002).
Although few have found recombinant DNA sequences in any organ or tissue sample from animals fed GM plants (EFSA, 2007), our results showed that small DNA fragments can also be detected in kids organs when mothers are fed GM soybean. Recombinant plant DNA has also been found by other authors as a rare event (Chowdhury et al., Reference Chowdhury, Kuribara, Hino, Sultana, Mikami, Shimada, Guruge, Saito and Nakajiama2003a; Mazza et al., Reference Mazza, Soave, Morlacchini, Piva and Marocco2005; Sharma et al., Reference Sharma, Damgaard, Alexander, Dugan, Aalhus, Stanford and McAllister2006) in organs and tissues of animals fed GM feeds. On the contrary, studies aiming at verifying the presence of transgenic fragments in milk gave negative results. In dairy cows fed GM soybean the chloroplastid sequence was found in leucocytes but not in milk (Klotz and Einspanier, Reference Klotz and Einspainer1998). In other trials carried out with dairy cows fed GM soybean meal and maize (Phipps et al., Reference Phipps, Deaville and Maddison2003; Nemeth et al., Reference Nemeth, Wurz, Artim, Charlton, Dana, Glenn, Hunst, Jennings, Shilito and Song2004) or goats fed GM maize (Rizzi et al., Reference Rizzi, Brusetti, Arioli, Nielsen, Tamagnini, Tamburini, Sorlini and Daffonchio2008), the chloroplast DNA fragments were detected in milk, but no transgenic fragments were found.
Detection of GM sequence in animal organs and tissues alone cannot justify the public concern regarding the human consumption of products from farm animals fed transgenic crops. Indeed, the implications of DNA transfer from food containing GM plants to an organism is no higher as compared with DNA transfer from food containing the corresponding conventional plant (Mazza et al., Reference Mazza, Soave, Morlacchini, Piva and Marocco2005).
Enzyme assay
In goats, a significant decrease in AST (day 60) and ALT (days 30 and 60) was detected in serum from goats fed GM soybean, but in both groups enzyme levels were in the normal range. Therefore, such difference should have no clinical relevance. Similarly, lower levels of ALT were found in serum from buffaloes fed transgenic cottonseed and the authors concluded that there were no differences in buffalo health status as determined by haematobiochemical constituents (Singh et al., Reference Singh, Tiwari, Kumar and Ravi Kumar2003). On all other days, and also in kids, the levels of the enzyme tested in serum did not show significant differences, thus suggesting that no adverse effects were induced by GM feed in treated animals, as previously reported in dairy cows (Yonemochi et al., Reference Yonemochi, Ikeda, Harada, Kusama and Hanazumi2003). By contrast, analysis of enzyme total and relative activities in tissues gave a different picture. As depicted in Figure 5 significant differences in enzyme levels concerned the kidney (higher levels of LDH and GGT) as well as heart and skeletal muscle (higher levels of LDH) in kids drinking milk from treated animals. This seems to indicate that some alteration occurred in kids even if serum levels were not affected. Moreover, LDH significantly increased in three organs (kidney, heart and skeletal muscle), suggesting that enzyme synthesis is generally altered in kids fed milk from GM-fed dams. The relative distribution of LDH isoenzymes confirms this hypothesis, showing significant differences in heart and kidney LDH1 and in muscle LDH1 and LDH2. In addition, if we compare the distribution of isoenzymes in percentage terms, LDH1 significantly increased also in the liver and heart LDH2 and LDH3 significantly decreased; a similar result was also seen for muscle LDH5. Therefore, the significant increase in LDH1 was accompanied by a percent decrease of other isoenzymes thus suggesting that, since LDH is a tetrameric enzyme made up of M and H subunits, a different combination of such subunits occurred in the heart (LDH-H4 > LDH-H3M1 and LDH-H2M2) and in the skeletal muscle (LDH-H4 and LDH3M1 > LDH-M4). A similar shift has been previously described in rabbits (LDH-H4 > LDH-H1M3) where the higher amount of H subunit was assumed to mean a higher specificity for the reduction of alpha-hydroxybutyrate to alpha-oxobutyrate, thereby showing that some metabolic changes occurred in the liver (Tudisco et al., Reference Tudisco, Lombardi, Bovera, D’Angelo, Cutrignelli, Mastellone, Terzi, Avallone and Infascelli2006b). Our data confirm this hypothesis: the isoenzyme mainly composed by the H subunit showed an increased percentage, which was also found in the muscle where LDH1 is not the dominant isoenzyme. Therefore, increased activity of LDH1 occurred in organs of GM-fed kids, as confirmed by histochemistry which showed a higher expression of total LDH in heart, kidney and skeletal muscles.
Since LDH1 is known to be involved in cell metabolism by favouring the reaction of lactate to pyruvate (Van Hall, Reference Van Hall2000), our results could indicate a general increase in cell metabolism. This hypothesis is in agreement not only with results obtained in rabbits but also with those who found significant modifications of some nuclear features in GM-fed mice suggesting a high metabolic rate and intense molecular trafficking (Malatesta et al., Reference Malatesta, Caporaloni, Gavaudan, Rocchi, Serafini, Tiberi and Gazzanelli2002). In any event, there is no scientific evidence that a local increase in LDH metabolism may be dangerous for health especially when it does not affect serum levels. Moreover, since serum activities of all the enzymes showed similar levels between the groups, it would be over speculative to assert that the GM diet was responsible for a local increase in LDH metabolism. However, the synthesis of LDH did actually change in more than one organ and such results should be taken into account for future research.
The mere detection of recombinant DNA fragments in animal organs and tissues cannot justify, by itself, public concerns regarding human consumption of products from farm animals fed transgenic crops. By contrast, although both BW and organ weight did not significantly differ between groups, any alteration in cell metabolism should be taken into account in this field. The modification in LDH synthesis, even without affecting serum levels of the enzyme, suggests an increase in cell metabolism. Therefore, possible long-term effects of such an alteration need to be elucidated.
In conclusion, our research attempted a wider survey in terms of detection of DNA fragments as well as investigating the impact of GM feed in terms of health. Our results showed that small DNA fragments can be detected in milk but also in kids organs when mothers are fed GM soybean and revealed elevated levels of LDH in tissues, suggesting an increase in cell metabolism. Although our data seem to conflict with the majority of studies in this field, they are consistent with other studies and the longer term consequences to health following GM food intake merit further consideration.















