Intermittent fasting (IF) results in many health benefits ranging from prevention to enhanced treatment of many metabolic diseases(Reference Longo and Panda1). Currently, IF was categorised into whole day fasting, every other day fasting and time-restricted feeding (TRF). Correspondingly, TRF refers to time-restricted food consumption for a period of counted hours and allows daily fasting duration greater than 12 h, irrespective of altering the quality and quantity of nutrients(Reference Engin2). In the context of therapeutic and sustained benefits of TRF, translational research is necessary to develop fasting-associated interventions into accessible, effective and inexpensive treatments with the potential to improve health span.
The circadian system is a 24-h daily rhythm in behaviour, physiology and metabolism that is continued under constant feeding–fasting, light–dark and sleep–awakening conditions(Reference Longo and Panda1). Agitation of circadian oscillator components leads to diabetes and obesity and develops impaired glucose tolerance and signs of metabolic diseases either in tissue-specific or whole-body loss of function in mouse models(Reference Bass and Takahashi3). The fasting cycle profoundly affects the circadian system by driving daily rhythm in the activities of nutrient homoeostasis key regulators including AMP-activated protein kinase (AMPK), cAMP response element-binding protein and protein kinase B(Reference Vollmers, Gill and DiTacchio4). AMPK degrades cryptochrome by promoting its phosphorylation(Reference Lamia, Sachdeva and DiTacchio5). Moreover, fluctuation in the level or activity of sirtuins especially sirtuin-1 (Sirt1) with the cellular energy state also disturbs the circadian system(Reference Masri and Sassone-Corsi6). SIRT1 activates the circadian system through recruitment to the circadian locomotor output cycles protein kaput (CLOCK):brain and muscle ARNT-like 1 (BMAL1) chromatin complex and by physically associating with CLOCK, to operate the circadian system(Reference Nakahata, Kaluzova and Grimaldi7), but there is lack of data on the effect of TRF on Sirt1 relating to gut microbiota. Although fasting-driven other metabolic regulators lead to coordinated oscillations at the transcriptional level(Reference Hatori, Vollmers and Zarrinpar8), which in turn synchronise daily circadian rhythms in cell repair, division, cellular metabolism and growth in a tissue-specific manner(Reference Mohawk, Green and Takahashi9).
Modern humans face complex health challenges in treating metabolic diseases. IF can be a therapeutic lifestyle approach for reducing the risk of many metabolic diseases like obesity and hypertension(Reference Hatori, Vollmers and Zarrinpar8). After consumption of 2092 kJ (500 kcal) with relatively high-protein diet for 2 d/week for 6 months reduced abdominal fat and blood pressure, while increased insulin sensitivity in overweight individuals(Reference Harvie, Pegington and Mattson10). IF may also diminish inflammation, as reported that practicing 2 months every other day fasting resulted in a significant reduction in inflammatory markers in asthma patients(Reference Johnson, Summer and Cutler11). TRF showed significantly improved rhythms and increased thermogenesis, which leads to reduced liver steatosis, adiposity, reduced serum cholesterol, normal glucose tolerance and increased bile acid production in mice(Reference Hatori, Vollmers and Zarrinpar8).
There is an additional link of nutrition and metabolic diseases with gut microbiome. Diet is a critical determinant of the gut microbial composition. Gut commensal bacteria and their metabolites can be both beneficial and harmful effects on host metabolism. Bacteria derived ammonia, hydrogen sulphide and N-nitroso compounds, from dietary protein, can activate inflammatory pathways and induce DNA damage and reactive oxygen species(Reference Kim, Coelho and Blachier12). In addition, the end product of dietary choline produced in the gut called trimethylamine-N-oxide correlates with CVD and stroke and promotes arteriosclerosis(Reference Tang, Wang and Levison13). IF confers protection in multiple sclerosis by increasing gut microbiome richness, changing their composition and metabolic pathway(Reference Cignarella, Cantoni and Ghezzi14). It is expected that TRF may trigger the fasting physiology within the time restriction paradigm. In turn, this process coordinates with robust circadian rhythm and enhances normal host metabolism in relation to gut microbiota.
Recently, IF is one of the attractive lifestyle intervention therapies for many metabolic diseases, although the underlying mechanism by which IF regulates circadian rhythm by activation of Sirt1 and modulates gut microbiota is unclear. Therefore, the present study for the very first time demonstrated that TRF modulates the circadian system through stimulation of Sirt1 and increases gut microbial diversity in human individuals. We found that microbiome richness and abundance are positively or negatively associated with circadian rhythm, lipid and liver profiles, respectively, indicating an important role of the microbiome as an integrator of the effects of TRF.
Methods
Participants and study design
In April 2018, we initiated a multi-centre study to examine the feasibility and effects of TRF on biomarkers and genetic predisposition of metabolic disease risks in young male healthy adults. This trial was conducted according to the guidelines laid down in the Declaration of Helsinki and approved by the ethical review committee of Nanjing Medical University, Jiangsu, China, under approval number (NJMUIRB (2018)004). International male students were recruited from three universities of Nanjing city, that is, Nanjing Medical University, Nanjing Agriculture University and Nanjing University of Science and Technology China. All participants provided a written consent form. Inclusion criteria for enrolment in the study were: (1) only male adults; (2) young aged; (3) absence of any ongoing chronic or infectious diseases; and (4) currently not using any supplements. Individuals were excluded if followed any strict diet plan or food for weight loss or gain. Females were excluded due to reproductive periods like menses as it influences the internal body biochemistry for short time. Participants were divided into the non-TRF group (n 24), in which participants received their regular diet on daily basis with no time restriction, and the TRF group (n 56), who underwent daily fasting of 16 h for 25 d. Study participants, assistant dietitian and the study principal investigator knew the group assignment after randomisation. The laboratory assistants and nurses who analysed and collected the samples were blinded to the group assignment.
Study procedures and sample collection
After enrolment, an assistant dietitian instructed the TRF group to strictly follow the study protocol. In the TRF group, participants were allowed to consume their normal diet with no food restriction for only 8 h per 24 h, that is, from 19.30 to 03.30 hours for 25 d. The assistant dietitian kept telephone and email contact with the participants to provide additional information as required and encouragement between the two visits. The non-TRF group continued their regular diet and were not given any specific instructions or time restriction. In the TRF group, the participants provided blood at pre-TRF (baseline) and post-TRF (after 25 d) and stool samples at one point, while from non-TRF, blood and stool samples were collected only one time after 25 d of trial at Sir Run Run Hospital affiliated with Nanjing Medical University. After completion of trial period, both groups (TRF and non-TRF) were informed through Wechat at the same time that not to take any food from 03.00 to 15.00 hours. The blood was drawn in the same time period between 15.00 and 18.00 hours from both groups. At each visit, the blood was drawn in the afternoon from both groups after a 12-h fast. Socio-demographic and body composition including body weight, BMI, %fat, waist to hip ratio and BMR were also measured as shown in online Supplementary Table S1.
Blood serum analysis
Venous blood samples were centrifuged at 4°C for 15 min at 4000 rpm. After collection, serum was portioned and sealed tubes were stored at –80°C until analysis. Serum concentrations of total cholesterol, LDL-cholesterol and HDL-cholesterol, TAG, alanine aminotransferase, aspartate aminotransferase and γ-glutamyl transferase activities were measured at laboratory of School of Public Health, Nanjing Medical University using a commercial reagent (Nanjing Jiancheng Bioengineering Institute) with the Roche/Hitachi Cobas c311 analyser (Roche Diagnostics) followed by previously described procedure(Reference Kowalska, Sciskalska and Bizon15,Reference Zeb, Safdar and Fatima16) .
ELISA
An ELISA kit (Youxuan Biotechnologies) was used to detect the serum level of IL-1β and TNF-α. Human serum samples were seeded in the microplates and then incubated with horseradish peroxidase-conjugate reagent for 1 h at 37°C. Chromogen solutions A and B were added for another 15 min at 37°C. Optical density values were measured at 450 nm within 15 min.
RNA extraction and real time-PCR
Total RNA was isolated from 200 µl serum samples using RNAiso Plus (TaKaRaBio Technology) and was reverse-transcribed to cDNA using the Prime Script TM RT Master Mix. The quantification of RNA was analysed by quantitative PCR using SYBR Premix Ex Taq II (TaKaRaBio Technology). The primer sequences are presented in Table 1. Clock, Bmal1, Sirt1, IL-1β and TNF-α primers and human U6 primers as an internal reference were obtained from RiboBio. All the PCR analyses were conducted using the 2−ΔΔCT method.
Table 1. Quantitative PCR primers

Clock, circadian locomotor output cycles protein kaput; Bmal1, brain and muscle ARNT-like 1; Sirt1, sirtuin-1.
Gut microbiota analysis
After sample collection from both groups, stools were immediately frozen at –80°C until DNA extraction. Metagenomic DNA was extracted using the Power Soil DNA Isolation Kit (Mo Bio Laboratories) according to manufacturer’s instruction. The V1–V3 region of the 16S rRNA gene was sequenced at Illumina Miseq v3 platform (2 × 300 bp paired-end reads) to comprehensively catalogue composition and abundance of the bacteria in the stool samples. 16S gene sequences were subjected to primer trimming, assembly by FLASH with default parameters(Reference Magoc and Salzberg17) and taxonomic assignment by Ribosomal Database Project classifier(Reference Wang, Garrity and Tiedje18). Diversity and richness in gut microbiota were measured as described(Reference Zhou, Gao and Mihindukulasuriya19).
Quantification and statistical analysis
Nutrient intake of the participants of both groups (TRF and non-TRF) is presented (as mean values and standard deviations) in online Supplementary Table S2. Differences in these characteristics between the two groups were compared using t test or the Mann–Whitney U test, as appropriate. The treatment effect for laboratory measures was evaluated using an ANCOVA. The ANCOVA model is a recommended approach for analysing baseline and follow-up measurements(Reference Vickers and Altman20). Analyses were performed using Graph Pad Prism 8.0. The abundance of a taxon in a stool sample was indicated as the relative abundance, which was calculated by dividing the number of reads for a taxon by the total read counts of the sample. Principal component analysis is an ordination technique that aims to discover the data pattern in N-dimensional spaces. It represents the major variation among objects in a reduced dimensional space. It is used to explore the microbiome data structure on the basis of dissimilarity measurement (Bray–Curtis dissimilarity) among samples(Reference Ramette21). Permutational multivariate ANOVA was used for formal statistical testing to investigate the whole microbial community difference between groups. ANCOVA was conducted only for those taxa with average relative abundance >0·05 %. Pearson or Spearman correlation was conducted to correlate log-transformed relative abundance of gut microbiome. The statistical analysis was performed using R (https://cran.r-project.org/). Differential analysis of the gut microbiome composition between two groups was performed using LEfSe, software to support high-dimensional class comparisons for microbiome data(Reference Segata, Izard and Waldron22). A false discovery rate q value was considered significant in all of the microbiome data analysis.
Results
Regulation of lipid profile by time-restricted feeding
Dyslipidaemia accounts one of the major risk factors of obesity, atherosclerosis and metabolic disorders. TRF reverses and inhibits diet-induced obesity and associated metabolic diseases in mice without changes in nutritional intake by altering the lipid levels(Reference Chaix and Zarrinpar23). For the first time, we investigated the antihyperlipidaemic effect of TRF in humans. Serum total cholesterol (P < 0·0001) and TAG (P = 0·0003 and P = 0·0052) levels reduced significantly in post-TRF compared with pre-and non-TRF, respectively (Fig. 1(a) and (b)), while LDL-cholesterol was remained constant in all groups (Fig. 1(c)). After TRF intervention, there was significant (P < 0·0001) increment in HDL-cholesterol level compared with non-TRF (Fig. 1(d)).
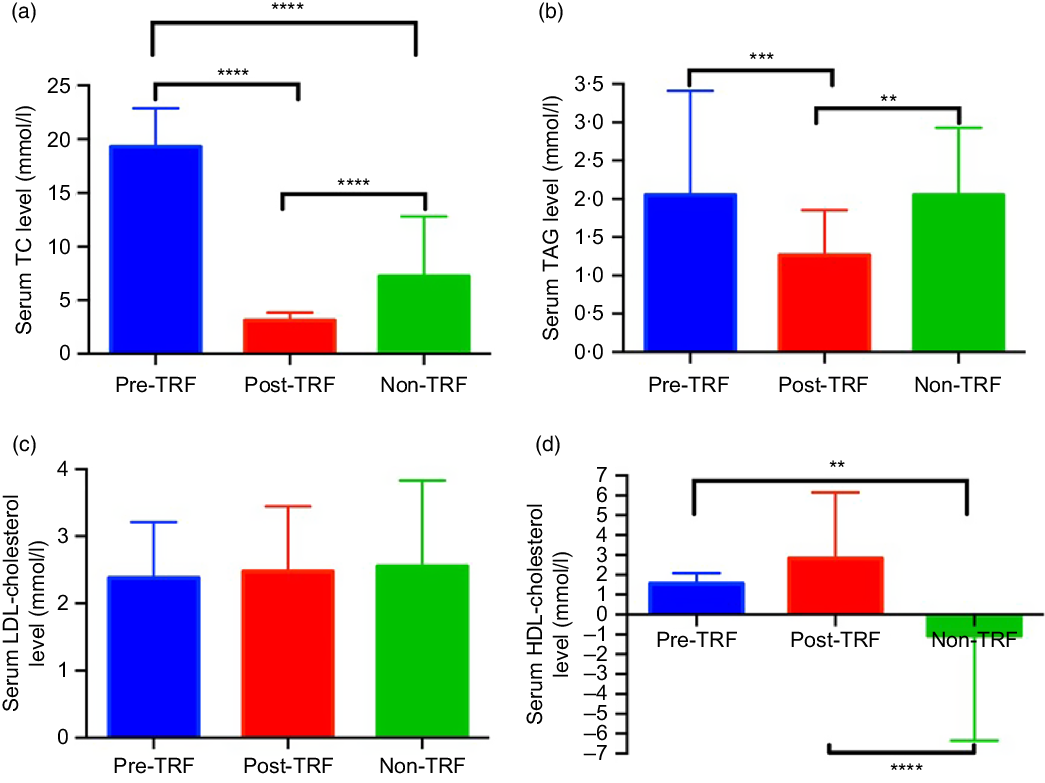
Fig. 1. Time-restricted feeding (TRF) reduces the risk of metabolic diseases through regulation of lipid profile. (a) Serum total cholesterol (TC), (b) TAG, (c) LDL-cholesterol and (d) HDL-cholesterol in pre/post-TRF (n 56) and non-TRF (n 24). Values are means, with standard deviations represented by vertical bars. One-way ANOVA followed by Bonferroni post hoc tests was used. **** P < 0·0000, *** P < 0·0001, ** P < 0·001 post-TRF v. pre-TRF and non-TRF.
Time-restricted feeding improves liver function by regulating its enzymes
Elevated liver enzymes especially alanine aminotransferase are associated with obesity-induced liver steatosis in mice fed a high-fat diet(Reference Nakahata, Kaluzova and Grimaldi7). Frequent and high energetic intake throughout the day and night exclusively attributed to obesity and ultimately disturb the liver enzymes and albumin level. In our results, alkaline phosphatase/γ-glutamyl transferase level was significantly (P < 0·0009) decreased in post-TRF than non-TRF (Fig. 2(a)). Serum aspartate aminotransferase (P = 0·0390 and P = 0·0268) and alanine aminotransferase (P = 0·0003 and P = 0·0174) were also reduced significantly in post-TRF compared with pre and non-TRF, respectively (Fig. 2(b) and (c)). TRF also has a profound effect on serum albumin, which recorded significantly (P < 0·0001) lower compared with pre-TRF (Fig. 2(d)). This indicates that TRF can regulate normal liver metabolism by modulating hepatic enzymes.

Fig. 2. Time-restricted feeding (TRF) improves liver function by regulating its enzymes. (a) Serum alkaline phosphatase (AKP), (b) aspartate aminotransferase (AST), (c) alanine aminotransferase (ALT) and (d) albumin in pre/post-TRF (n 56) and non-TRF (n 24). Values are means, with standard deviations represented by vertical bars. One-way ANOVA followed by Bonferroni post hoc tests was used. **** P < 0·0000, *** P < 0·0001, * P < 0·01 post-TRF v. pre-TRF and non-TRF.
Time-restricted feeding reduced production of pro-inflammatory cytokines
Excess nutrients intake induces inflammatory response, which has been causally linked to dysregulation of glucose and lipid metabolism(Reference Gregor and Hotamisligil24). Dyslipidaemia and obesity-associated inflammation predispose to the pathogenesis of atherosclerosis, which is a clinical manifestation of vascular inflammation in metabolic diseases(Reference Rocha and Libby25). Regarding the role of inflammation in atherosclerosis, IL-1β is increased in atherosclerosis and associated with disease severity(Reference Masters, Latz and O’Neill26). TNF-α and IL-1β are the most attributable pro-inflammatory cytokines of metabolic dysregulation secreted by adipose tissue(Reference McArdle, Finucane and Connaughton27). Therefore, we studied the effect of TRF on IL-1β and TNF-α. However, the mRNA and serum levels of both cytokines down-regulated in the post-TRF group compared with pre- and non-TRF groups but did not reach statistical significance (Fig. 3(a)–(d)).
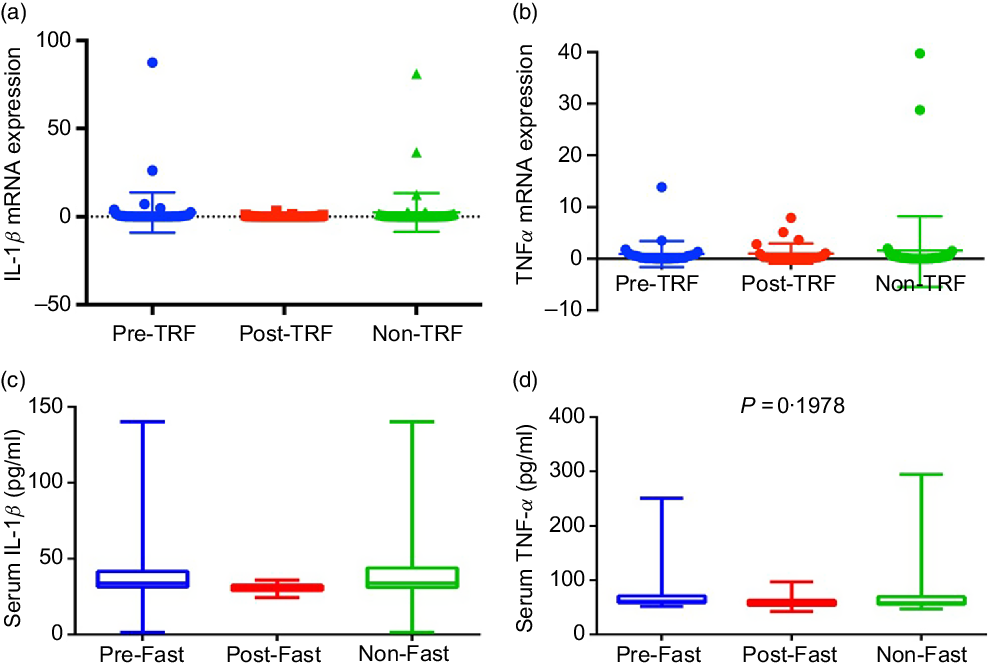
Fig. 3. Time-restricted feeding (TRF) reduces production of pro-inflammatory cytokines. (a) IL-1β and (b) TNF-α gene expression at mRNA and (c and d) serum levels in pre-TRF, post-TRF (n 56) and non-TRF (n 24). One-way ANOVA followed by Bonferroni post hoc tests was used.
Time-restricted feeding regulates circadian rhythm and its stimulator
Feeding- and fasting-driven metabolic regulators were associated extensively with circadian oscillation that coordinated in a number of metabolic pathways(Reference Nakahata, Kaluzova and Grimaldi7). However, under nutritional challenges such as one posed by high-fat diet, circadian oscillations are necessary and sufficient for preventing metabolic disorders(Reference Yang, Kang and Liu28). Therefore, we have identified in humans that TRF improves the circadian oscillation for normal metabolic regulation. The mRNA level of Bmal1 (P = 0·0020) and Clock (P = 0·0302) genes was significantly up-regulated by TRF intervention compared with the pre- and non-TRF groups, respectively (Fig. 4(a) and (b)). Sirt1 activation modulates the circadian physiology in mice(Reference Bellet, Zocchi and Sassone-Corsi29). In the present study, we showed that activation of Sirt1 can regulate the circadian rhythm. As in the post-TRF group, the mRNA level of Sirt1 was significantly (P = 0·0068 and P = 0·0300) up-regulated compared with the pre- and non-TRF groups, respectively (Fig. 4(c)).
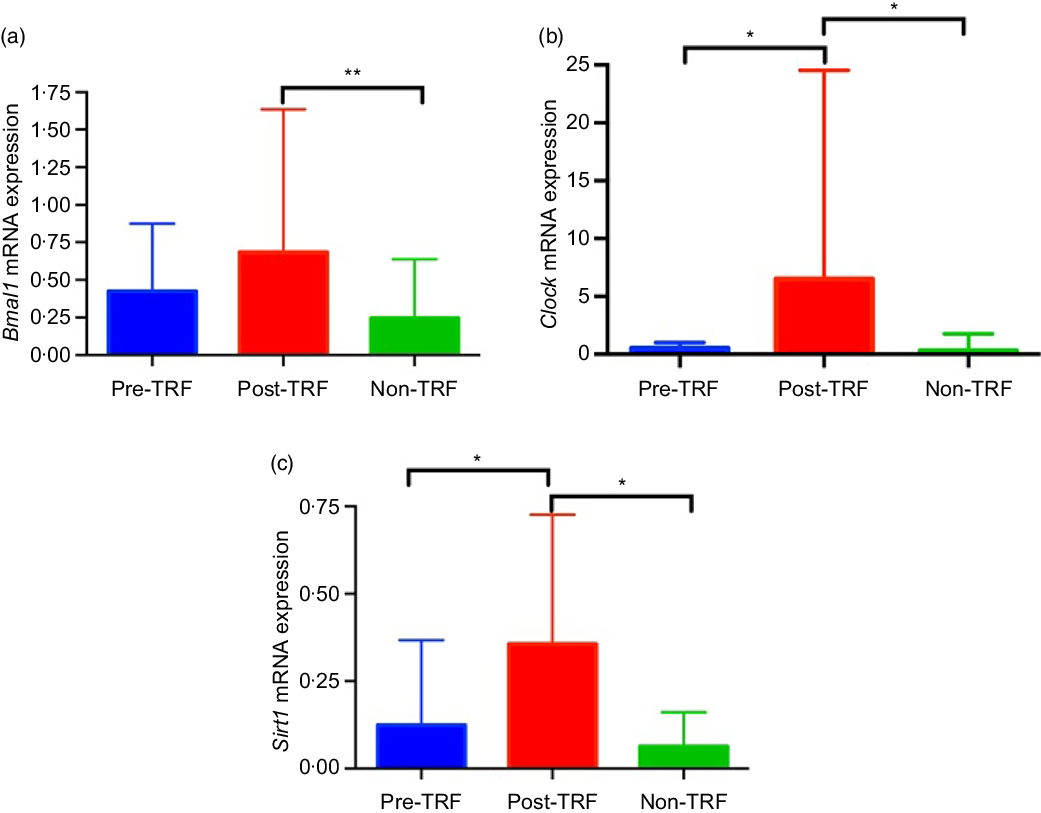
Fig. 4. Time-restricted feeding (TRF) regulates circadian rhythm and its stimulator. (a) Brain and muscle ARNT-like 1 (Bmal1), (b) circadian locomotor output cycles protein kaput (Clock), (c) sirtuin-1 (Sirt1) expression at the mRNA level. Values are means, with standard deviations represented by vertical bars. One-way ANOVA followed by Bonferroni post hoc tests was used. ** P < 0·005, * P < 0·05 post-TRF v. pre-TRF and non-TRF.
Time-restricted feeding increases gut microbiome diversity and has profound association with sirtuin-1 expression and HDL level
Recently, animal and human studies have discovered that reduced microbiome diversity, altered gut microbial activities and scattered microbiome relative abundance especially of two phyla, that is, Bacteroidetes and Firmicutes, are associated with obesity development(Reference Conterno, Fava and Viola30,Reference Weinberger, Schwartzstein and Weiss31) . Using the relative abundance of the differential abundant microbial taxa, we separated the post-TRF microbiota from the non-TRF group. However, there is no substantial overlap between the two groups as illustrated by applying principal component analysis (Fig. 5(a)). Microbial richness is a measure of α diversity for gut microbiota(Reference Lozupone and Knight32) and reflects complexity of a microbial community with higher diversity being associated with more healthy gut microbiome(Reference Heiman and Greenway33,Reference Le Chatelier, Nielsen and Qin34) . TRF significantly increased microbial diversity compared with the non-TRF group (Fig. 5(b), P < 0·005). Gut microbiota might modulate systemic metabolic responses(Reference Sonnenburg and Backhed35), and we also observed that TRF modulates gut microbiota, which exhibit regulation of genetic pathway. Indeed, the Sirt1 expression was positively correlated with gut microbiome richness (r 0·45, P < 0·0201, Pearson correlation; Fig. 5(c)). Serum HDL level showed a trend towards positive association (r 0·42, P < 0·0289, Pearson correlation; Fig. 5(d)), while serum albumin has significant negative correlation with gut microbiome richness (r –0·37, P < 0·0495; Fig. 5(e)). This indicates that TRF reduces the burden of metabolic risk through regulation of Sirt1 expression, serum HDL and albumin levels induced by gut microbiome modulation.
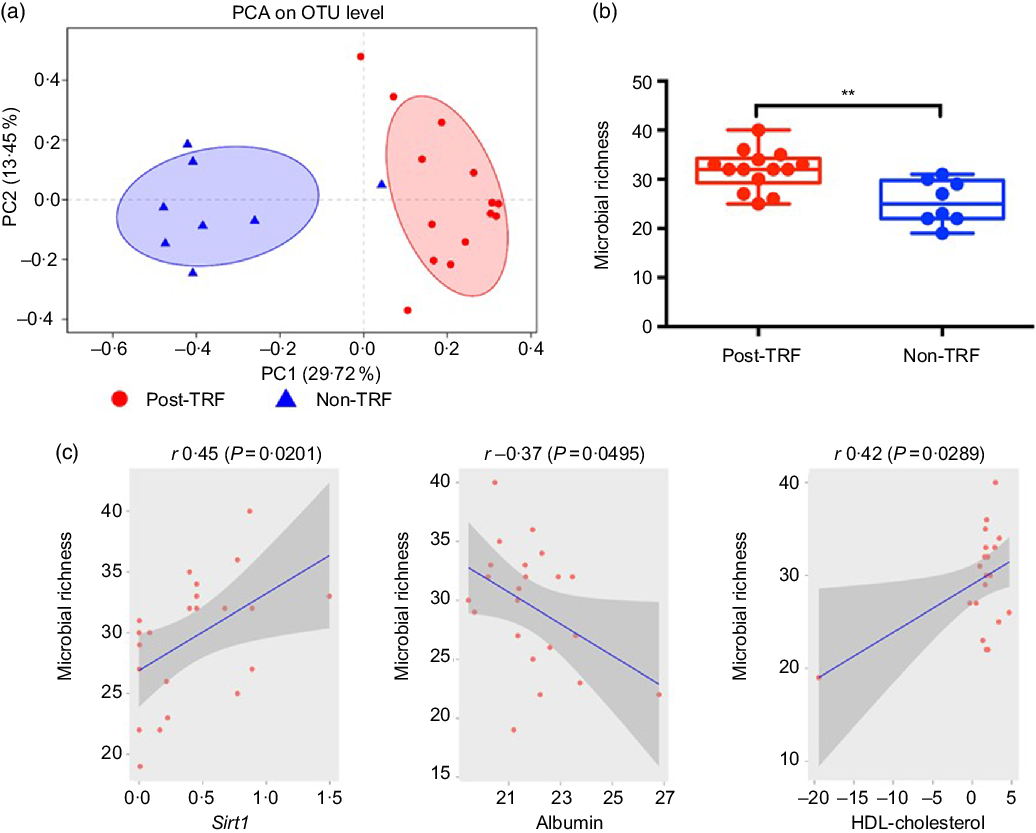
Fig. 5. Time-restricted feeding (TRF) increases gut microbiome diversity and has profound association with sirtuin-1 (Sirt1) expression and HDL level. Stool samples were collected from the post-TRF (after 25 d of trial, n 14) and non-TRF (n 18) groups. (a) Principal component analysis (PCA) illustrates microbiome similarity in the post-TRF and non-TRF groups at the operational taxonomic unit (OTU) level (each dot is one sample; x axis and y axis are first and second dimensions of microbiome data, respectively). Samples from two groups clustered separately, indicating two distinct microbiome communities (P < 0·05, permutational multivariate ANOVA test). (b) Microbial richness increased significantly in post-TRF but not in non-TRF (linear regression ** P < 0·005). (c) Sirt1 mRNA expression was significantly positively correlated with microbial richness (r 0·45; P = 0·0201, Pearson correlation). (d) Serum albumin was significantly negatively correlated with microbial richness (r −0·37, P = 0·0495). (e) Serum HDL level was significantly positively correlated with bacterial richness (r 0·42, P = 0·0289).
Time-restricted feeding results in significantly different microbial compositions in gut microbiome
We applied linear discriminant analysis combined effect size measurements to explore the significant changes and relative richness of bacterial community in both groups. At genus level, 34 and 18 bacteria were enriched in the TRF and non-TRF groups, respectively. Prevotellaceae (prevotella_9 and prevotella_2) and Bacteroidetes were the most abundant in the TRF group with greater than 2·5 linear discriminant analysis score, while in the non-TRF group, Escherichia, Shigella and Peptostreptococcus were abundant at the genus level (Fig. 6(a)). Further, we found an association of LDL-cholesterol and TAG with Bacteroidia belonging to the phylum Bacteroidetes that maintain a complex and generally beneficial relationship with the host when retained in the gut(Reference Wexler36). The relative richness of Bacteroidia has a significant negative association with LDL-cholesterol (r –0·46, P = 0·0186) and TAG (r –0·40, 0·0364) (Fig. 6(b) and (c)). This indicates that the reduced serum level of LDL-cholesterol and TAG are probably related with increased Bacteroidia diversity.

Fig. 6. Time-restricted feeding (TRF) results in significantly different microbial compositions in the gut microbiome. 16S rRNA sequencing was performed for stool collected from post-TRF (n 18) and non-TRF groups (n 14). (a) Linear discriminate analysis (LDA) was conducted to identify differentially represented microbial communities abundance in the two groups. Microbial community abundance with >2·5 LDA score (x axis) and P < 0·05 is shown. The right side of the figure represents microbial community whose abundance was significantly higher in the post-TRF group, while the left side in the non-TRF group. The absolute LDA value is the effect size between two groups for a particular microbial community. (b and c) Serum LDL-cholesterol and TAG have significantly negative correlation with Bacteroidia.
Association of healthy gut microbiome with circadian genes
After the determination of combined microbial richness, we further analysed whether there is any association of specific microbiome with circadian genes. We found that some of the beneficial bacteria including Prevotellaceae (r 0·67, P = 0·0003), prevotella_9 (r 0·70, P = 0·0002) and Bacteroidia (r 0·40, P = 0·0370) have a significant positive association with Bmal gene (Fig. 7(a)–(c)). Previous studies suggested that Sirt1 functions as a molecular link between circadian rhythms and metabolic control and also an important modulator and activator of Clock gene expression(Reference Bellet, Nakahata and Boudjelal37,Reference Liu38) . Interestingly, we reported that the expression of Clock and Bmal1 probably up-regulated with the increase in Sirt1 expression. Here, the relative abundance of Prevotellaceae (r 0·70, P = 0·0002), Prevotella_9 (r 0·73, P < 0·0001), Bacteroidia (r 0·45, P = 0·0199), Dialister (r 0·50, P = 0·0105) and Prevotella_2 (r 0·69, P = 0·0002) was significantly positively associated with Sirt1 expression (Fig. 7(d)–(h)).
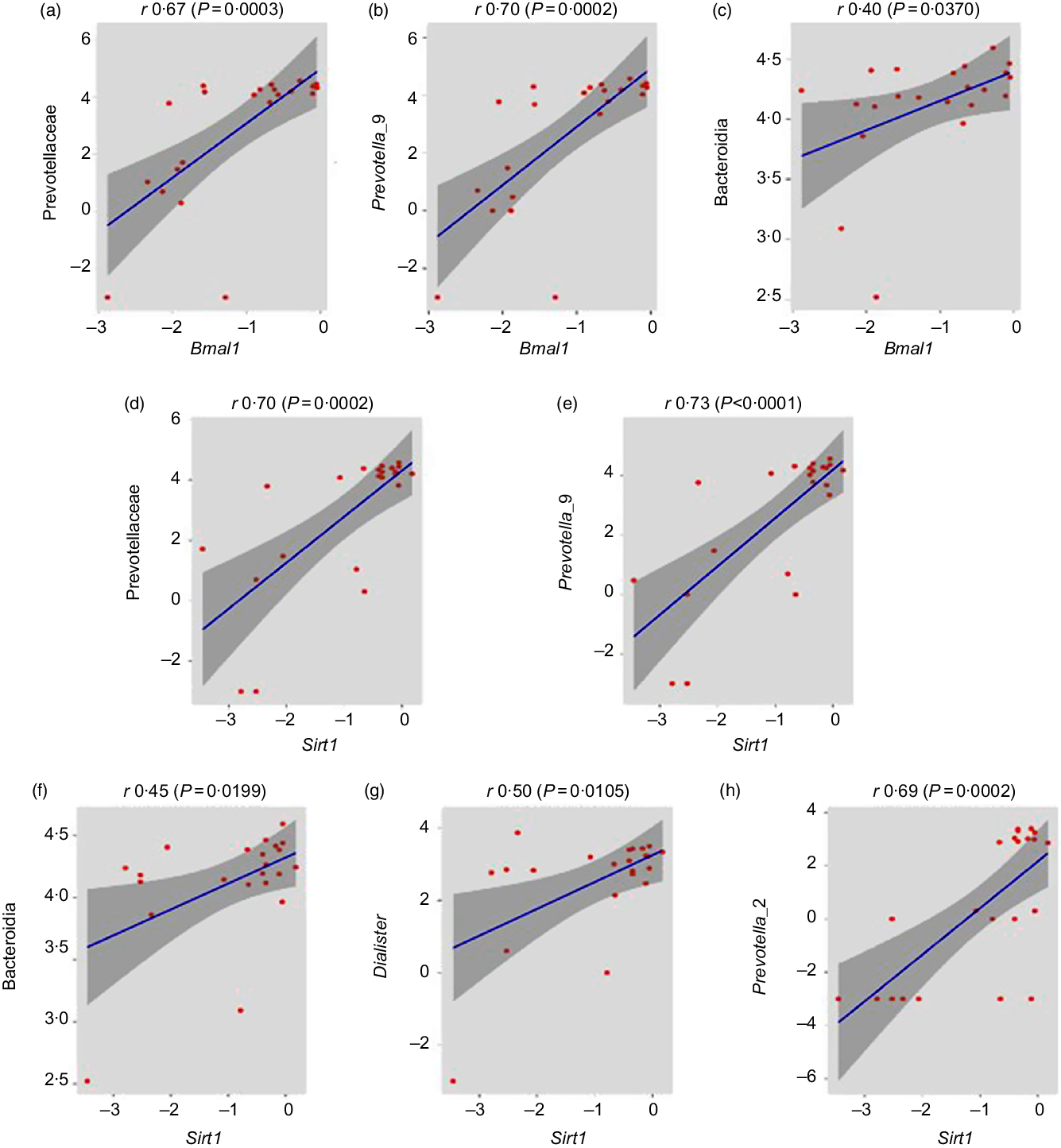
Fig. 7. Association of healthy gut microbiome with circadian genes. (a–c) Prevotellaceae (family), Prevotella_9 (genus) and Bacteroidia (class) abundances were significantly positively correlated with brain and muscle ARNT-like 1 (Bmal1) gene expression (r 0·67, P = 0·0003; r 0·70, P = 0·0002; r 0·40, P = 0·0370, respectively, Pearson correlation). (d–h) Prevotellaceae (family), Prevotella_9 (genus), Bacteroidia (class), Dialister (genus) and Prevotella_2 (genus) abundance have significantly positive association with sirtuin-1 (Sirt1) gene expression (r 0·70, P = 0·0002; r 0·73, P < 0·0001; r 0·45, P = 0·0199; r 0·50, P = 0·0105 and r 0·69, P = 0·0002, respectively).
Discussion
Disturbed dietary/feeding pattern bothers metabolic pathways entrained by both feeding and circadian rhythms. This chronological misalignment in cellular metabolism in combination with nutrient quality and gut microbiota predisposes obesity and the metabolic syndrome(Reference Longo and Panda1). Many studies have observed that IF exerts a strong anti-obesity, anti-diabetic, anti-inflammatory and cardioprotective effects in animal models(Reference Bass and Takahashi3). Currently, the recommended lifestyle modification focused on the alteration of individual nutrition and gut microbiome that are interlinked with metabolic risk. This is the first report that found that the mRNA expressions of Sirt1 and circadian genes (Bmal1 and Clock) were higher and the microbiota were improved after TRF, the beneficial effect of TRF probably related to the Sirt1 activation. Therefore, we introduced TRF, a lifestyle intervention, that can prevent metabolic risk through modification of circadian rhythm and gut microbiome by preserving natural feeding rhythms.
Lipid homoeostasis was maintained by the interaction of circadian components with metabolic regulators. Herein, we show that TRF reduced the lipid level including total cholesterol and TAG and increased HDL in the circulation. Similarly in a previous study, intermittent energy restriction induced a decrease in total cholesterol and TAG(Reference Harvie, Pegington and Mattson10). TRF reduced fatty acid synthesis and increased fat oxidation, which in turn leads to reduction in adiposity(Reference Chaix and Zarrinpar23), and it is demonstrated that TRF is a lifestyle therapy against metabolic diseases by inhibiting dyslipidaemia(Reference Hatori, Vollmers and Zarrinpar8).
Liver metabolism and homoeostasis were maintained by mutual coordination of metabolic regulatory enzymes and circadian oscillation. Altered liver metabolism induced by increased level of hepatic enzymes could be a risk for the development of metabolic diseases. We showed that TRF reduced the serum level of liver enzymes including alkaline phosphatase, aspartate aminotransferase, alanine aminotransferase and albumin, as reported previously in mice that TRF prevents hepatomegaly and liver failure by reducing the serum alanine aminotransferase(Reference Hatori, Vollmers and Zarrinpar8).
In animal models, IF has shown potential role against inflammation(Reference Arumugam, Phillips and Cheng39). Therefore, we studied the effect of TRF on IL-1β and TNF-α in humans. The reduction in these two cytokines was non-significant. Conversely, fasting mimicking diet improved inflammation by returning C-reactive protein, a marker of inflammation in CVD in a very limited sample of seven subjects(Reference Brandhorst, Choi and Wei40). IF also reduced inflammation through less production of IL-17 by altering gut microbiota in mice with multiple sclerosis(Reference Cignarella, Cantoni and Ghezzi14). However, every other day fasting significantly reduced TNF-α in patients suffering from asthma after 2 months(Reference Magoc and Salzberg17). These discrepancies may be due to sample size or mode of fasting, inflammation markers and even study subjects.
Food restriction not only increases the rhythmic transcripts but also improves amplitude, synchronises the phases of oscillations and reduces food intake. Feeding affects the clock through nutrient sensing pathway by anabolic targets. The mTOR pathway phosphorylates casein kinase 1 which further phosphorylates the clock component called PER, as a result changing its stability(Reference Zheng and Sehgal41). Besides, erratic dietary patterns including disturbed feeding, eating attitude and poor nutrition can cause chronic disruption in the circadian rhythm that increases metabolic risk. As previously reported that after 10 d of misalignment of circadian system, individuals developed insulin resistance, elevated post-prandial glucose and increased arterial pressure(Reference Scheer, Hilton and Mantzoros42). Improvement in circadian rhythm helps to prevent or reverse metabolic diseases. This is in line with a study reported that Sirt1 gene expression was increased after 48 h of fasting in ten healthy lean men(Reference Edgett, Scribbans and Raleigh43). In disease condition, the circadian genes usually down-regulated as reported previously, that Bmal1 and Sirt1 expression decreased significantly and resulted in aggravation of osteoarthritis-like gene expression changes under IL-1β stimulation. Overexpression of Bmal1 through stimulation of Sirt1 relieved the metabolic imbalance induced by IL-1β in osteoarthritis(Reference Edgett, Scribbans and Raleigh43). Therefore, we can suggest that Sirt1 probably acts as a stimulator for circadian genes.
In addition, TRF increased gut microbiome richness and abundance at different levels. Importantly, we found that Sirt1 activation and increased level of HDL-cholesterol were significantly positively associated with an increase in microbial richness. These data strongly suggest that TRF-induced alteration in gut microbiome richness can intervene a systemic anti-obesity response and protection against metabolic risk, which is in agreement with the previous study(Reference Cignarella, Cantoni and Ghezzi14). It is well known that nutrition plays an important role in gut microbiome modulation(Reference Muegge, Kuczynski and Knights44). Gut microbiota dysbiosis mediates the pathogenesis of several human diseases(Reference Guinane and Cotter45). In the present study, we found that TRF increased microbial diversity and was positively correlated with HDL and Sirt1. These results are in consistent with previous studies that reported that the gut microbial community structure was significantly influenced and more diverse after long-term intervention of CR than people eating Western diets(Reference Griffin, Ahern and Chen46), while IF also significantly increased the microbiome richness(Reference Cignarella, Cantoni and Ghezzi14). TRF had a striking effect on microbiome composition. Interestingly, we also showed that Bacteroidia inversely correlated with LDL-cholesterol and TAG level that exhibited anti-obesity response. This is in line with a study that suggested that increased abundance of Bacteroidetes members is directly associated with weight loss in mice(Reference Ridaura, Faith and Rey47). In our study, TRF exhibited a significant increase in the abundance of Bacteroidia and Bacteroidetes, which is in agreement with previous reports in animal models(Reference Kohl, Amaya and Passement48,Reference Queipo-Ortuno, Seoane and Murri49) . Low abundance of Bacteroidetes enhanced the development of inflammatory conditions including obesity, atherosclerosis, neurodegenerative diseases and diabetes(Reference Mouzaki, Comelli and Arendt50).
Dysbiosis and disturbed interaction of host-microbe may cause circadian misalignment leading to the pathogenesis of metabolic disorders. Overexpression of Bmal1 decreases hyperlipidaemia and atherosclerosis through modulation of lipoprotein production and biliary cholesterol excretion in ApoE−/− mice(Reference Pan, Bradfield and Hussain51). Besides food intake and daylight cycle, microbiota are also involved in control of circadian system that regulates intestinal physiology and systemic metabolism(Reference Huang, Pai and Yu52). Gut microbiome has circadian rhythmicity and helps the host circadian clock function. Here, we found a significant positive correlation of Bmal1 with Prevotella and Bacteroidia and Sirt1 with Prevotellaceae, Bacteroidia and Dialister. Prevotella-derived SCFA produced by the fermentation of non-digestible fibre facilitate peripheral clock adjustment in mouse tissues(Reference Tahara, Yamazaki and Sukigara53). Bmal1 expression disturbs during gut microbiome ablation resulting in prediabetic phenotype and ileal corticosterone overproduction. In addition, lacking healthy gut microbiota leads to a trend in down-regulation of clock control gene expression, especially involved in metabolic regulation(Reference Leone, Gibbons and Martinez54).
In conclusion, the present study showed that TRF reduced serum lipid and liver profiles, while it induced down-regulation of inflammatory genes. In addition, TRF increased microbial richness and abundance, which is probably associated with circadian rhythm and lipid levels. Therefore, TRF could be a safe therapeutic remedy for the prevention of metabolic diseases related to dyslipidaemia and elevated liver function, as it regulates circadian rhythm associated with gut microbiome modulation.
Acknowledgements
This work was supported by funding from the Priority Academic Program Development of Jiangsu Higher Education Institutions and China Scholarship Council (CSC).
The authors declare that there are no conflicts of interest.
Supplementary material
For supplementary material referred to in this article, please visit https://doi.org/10.1017/S0007114519003428











