INTRODUCTION
Shiga toxin-producing Escherichia coli (STEC) serotype O157:H7 emerged as a significant risk to public health in the 1980s, associated with outbreaks of gastrointestinal symptoms and haemolytic uraemic syndrome (HUS) [Reference Adams1]. Microbiological and epidemiological investigations identified ruminants (mainly cattle and sheep) as the zoonotic reservoir [Reference Chapman2]. Transmission occurs by consumption of contaminated food or water, or direct contact with animals or their environment [Reference Byrne3]. The infectious dose is low (10–100 organisms) and person-to-person spread can occur in households, nurseries and other institutional settings.
Prior to 2005, outbreaks in England were often linked to the consumption of contaminated undercooked beef or cross-contamination of cooked meats [Reference Adams1]. Since that time, reports of meat-related outbreaks have decreased and outbreaks associated with eating contaminated raw vegetables and salad leaves have been reported more regularly [Reference Launders4–Reference Sinclair6]. The largest national outbreak in the United Kingdom (252 cases) was caused by STEC O157:H7 PT8 occurred in 2011 and was linked to handling raw leeks and potatoes [Reference Launders7].
At Public Health England (PHE), whole genome sequencing (WGS) is used for the routine surveillance and detection of outbreaks caused by STEC O157:H7. WGS data have facilitated a more robust approach to case ascertainment and provided insight into the investigation of complex outbreaks scenarios [Reference Jenkins8–Reference Rowell10]. In August 2015, an outbreak of STEC O157:H7 phage type (PT) 8 was detected following routine monitoring of the WGS typing data. The aim of this study was to identify the original source and likely transmission routes associated with this outbreak.
METHODS
Microbiology
In England and Wales, all presumptive isolates of STEC O157:H7 from clinical cases detected at local hospital laboratories, and isolates from food, animal and environmental samples detected at commercial or government-funded laboratories, are referred to the Gastrointestinal Bacteria Reference Unit (GBRU) at PHE for confirmation and further typing. Isolates are confirmed as STEC O157 using PCR targeting stx1, stx2, eae encoding the adhesin, intimin and O157rfb which is part of the O157 antigen encoding cluster. All isolates submitted during the period of this study were phage typed and genome sequenced. Genes encoding the H7 flagella antigen were detected in the genome sequence, as previously described [Reference Joensen11].
For WGS, DNA was extracted from cultures of STEC O157:H7 for sequencing on the Illumina HiSeq 2500 instrument. High-quality Illumina reads were mapped to the STEC O157:H7 reference genome Sakai (Genbank accession BA000007) using BWA-MEM [Reference Li and Durbin12]. Single nucleotide polymorphisms (SNPs) were identified using GATK2 [Reference McKenna13] in unified genotyper mode. Core genome positions that had a high-quality SNP (>90% consensus, minimum depth 10×, GQ ⩾ 30) in at least one isolate were extracted and RaxML [Reference Stamatakis14] was used to derive the maximum likelihood phylogeny of the isolates. Isolates with <30 × coverage when mapping to the O157:H7 reference genome were discarded from analysis. Positions with <10 × coverage or where the consensus base was present in <90% of the reads were discarded from analysis. The average coverage obtained in the study was 55·5×.
Genomes were compared to the sequences held in the PHE STEC O157:H7 WGS database. Isolates of STEC O157:H7 with less than five SNPs differences within their core genome are considered closely related and likely to have an epidemiological link [Reference Dallman15]. Hierarchical single linkage clustering was performed on the pairwise SNP difference between all isolates at various distance thresholds (Δ250, Δ100, Δ50, Δ25, Δ10, Δ5, Δ0) [Reference Dallman16]. The result of the clustering is a SNP address that can be used to describe the population structure based on clonal groups. Although isolates greater than 5 SNPs apart are unlikely to be part of the same temporally linked outbreak, deeper phylogenetic relationships within the 10 or 25 SNP clusters may provide epidemiologically useful information or associations [Reference Dallman15]. Stx subtyping was performed as described by Ashton et al. [Reference Ashton17].
Timed phylogenies were constructed using BEAST-MCMC v1.80, as previously described [Reference Dallman15]. A relaxed log normal clock rate under a constant population size was selected as found to be optimal for STEC O157:H7 in our previous study [Reference Dallman18]. The model was run with a chain length of one billion and a maximum clade credibility tree was constructed using TreeAnnotator v1.75.
FASTQ reads from all sequences in this study and the PHE STEC O157:H7 WGS data can be found at the PHE Pathogens BioProject at the National Center for Biotechnology Information (Accession PRJNA248792) (Supplementary Table S1).
Epidemiological investigations
Presumptive cases of STEC are reported directly to PHE centres by clinical microbiologists at local hospital laboratories. A standardised STEC Enhanced Surveillance Questionnaire (SESQ) (https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/323423/VTEC_Questionnaire.pdf) is administered to cases either by local health protection professionals or environmental health practitioners (EHPs). Data from the questionnaires are entered into the National Enhanced Surveillance System for STEC in England (NESSS) [Reference Adams1].
Case–case investigations
Based in the initial analysis of the SESQ data, a case–case study design was used to test the hypothesis that outbreak cases were associated with prepacked salad from the national retailer (retailer X). The analysis was conducted in Stata 13.0. Existing STEC enhanced surveillance questionnaires were used for both outbreak cases and control cases. Only responses to binary questions were included in the analysis. Outbreak cases were defined as primary symptomatic cases of STEC O157:H7 PT8 stx2a, belonging to the same five SNP single linkage cluster (hereafter referred to as the outbreak strain), with the SNP address 18·35·380·765·1009·1526.% confirmed by GBRU with onset of diarrhoea between 19 July and 30 September 2015, and resident in England or Wales. Only the first 45 outbreak cases were included in this analysis, as the remaining three cases were identified after the case–case study had been completed. For each outbreak case, two control cases were selected from indigenous cases of STEC O157:H7 with PTs other than PT8 associated with illness in July and August reported to the national surveillance system during the preceding years (2010–2014). Outbreak cases and control cases were not matched on geography since cases were distributed across England. Odds ratios were calculated for each single binary exposure with a χ 2 test; multivariable logistic regression was then performed using a forward–backwards approach: first, exposures with a P value <0·05 were included and a backwards stepwise procedure undertaken. Exposures found to be significant or potential confounders were kept. The rest of the exposures (P value 0·05–0·2) were added to a second model and the same backwards stepwise procedure undertaken. An alternative model was also constructed in which control cases were restricted to those with onset in July and August 2015, to account for a change in the market share of retailer X in the preceding year.
A second case–case study was conducted to test the hypothesis that illness in cases detected after the outbreak, with isolates within 10 SNPs of the outbreak strain, was associated with consumption of lamb meat and/or exposure to live sheep or lambs. The analyses were conducted with the statistical software package R version 3.3.2 (http://www.R-project.org). Cluster cases were defined as primary or co-primary with isolates belonging to the 10 SNP single linkage cluster designated 18·35·380·765·1009.%, and onset between 1 January and 31 December 2016. Indigenous cases of STEC O157:H7 infection reported to the national surveillance system from 2009 to 2016 inclusive with PTs other than PT8, frequency-matched to cases in terms of month of onset and PHE centre of residence, were used as control cases. In this analysis, details from free text responses were also included and parsed into single word binary variables with natural language processing techniques (http://www.jstatsoft.org/v25/i05/) [Reference Feinerer19]. A high ratio of control cases to cluster cases was used (120:1; 1919 controls, 16 cases) to facilitate analysis of free text-derived variables with low representation. Odds ratios were calculated with Fisher's exact test. Multivariable logistic regression was conducted on exposures from the single variable analysis that were experienced by at least 20% of cases and had an odds ratio and lower 95% confidence interval >1. Age and sex were also included in the model a priori. A forward–backwards approach was used to derive the final model, as described in the previous section.
Microbiological examination of food and environmental samples
Food, water and environmental samples were collected by EHPs from the retailer (n = 68), distributor (n = 24) and the five growers implicated in the supply chain (n = 55) (Supplementary Table S2), and transported in accordance with the Food Standards Agency Food Law Code of Practice (https://www.food.gov.uk/enforcement/codes-of-practice/food-law-code-of-practice-2015) to PHE Food, Water and Environmental Microbiology Laboratories at Preston, Porton, London, York or Birmingham in cold boxes at a temperature of between 0 and 8 °C and tested within 24 h of collection. Testing of the food samples followed PHE Standard Method F17 based on BS EN ISO 16654:2001 http://img.21food.cn/img/biaozhun/20100729/181/11294219.pdf, as previously described [Reference Jenkins8].
Investigation of animal movements with the network software FoodChain-Lab
The locations and details of farms and lairage where lambs associated with the contaminated meat products were kept immediately prior to slaughter, and farms where cases had reported exposure to live sheep in the 7 days preceding the onset of symptoms were obtained and recorded. Data on the movement of sheep and lambs to and from these sites were requested from the Animal Reporting & Movement Service (ARAMS) (http://www.arams.co.uk/), a system which records the movements of sheep, goat and deer between agricultural premises, livestock markets and slaughterhouses in the UK. All movement data between May 2015 and October 2016 were requested. The locations and details of farms located most closely to the salad growers associated with the 2015 outbreak were also obtained and recorded but data for sheep and lamb movement onto and off these sites were not available for all requests for information.
Using the open source software, FoodChain-Lab (http://silebat.github.io/BfROpenLab/fcl_home.html), a network of sheep and lamb movements was constructed. FoodChain-Lab facilitates the identification of possible outbreak sources or vehicles through calculation of tracing scores for venues and food products under investigation [Reference Weiser20]. A premise linked to a confirmed case was weighted (assigned as 1 vs. 0 for holdings without confirmed cases). Tracing scores were calculated for all movements. Tracing scores are a value between 0 and 1. Higher tracing scores are correlated with the probability that an animal moved on or off a holding and may help to identify a common contamination source.
RESULTS
Epidemiological investigation of the outbreak associated with consumption of contaminated salad
On 11 August 2015, four cases of STEC O157:H7 stx2a PT8 belonging to the same five SNP single linkage cluster (18·35·380·765·1009·1526.%) were identified. By 31 October 2015, the cluster comprised 47 cases (Fig. 1). Onset dates for confirmed and probable cases ranged from 29 July to 30th September 2015 (Fig. 2). None of the cases developed HUS, but 16 (34%) were hospitalised and 34 cases (72%) had blood in their stools. The majority of cases were over 18 years of age (88%) and female (69%). Cases were dispersed throughout England, and there was one case in South Wales (Fig. 2). The highest burden of cases was in the North West, South East and East of England (nine cases each).
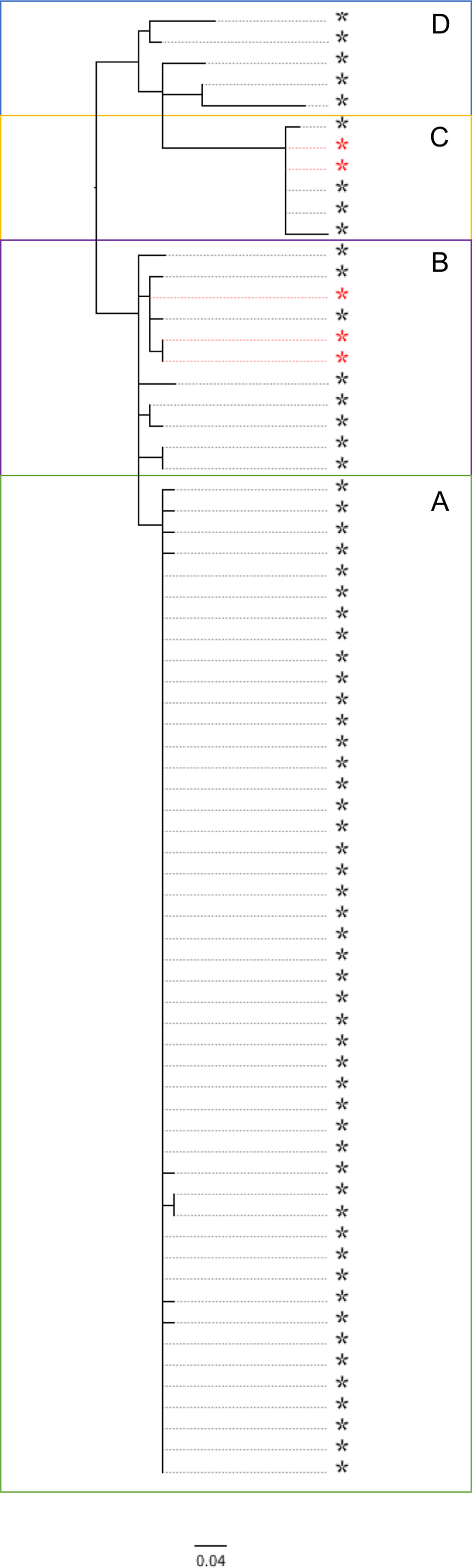

Fig. 1. Phylogenetic relationship between the isolates linked to the salad outbreak sub-cluster A (highlighted in green), and non-salad outbreak isolates (sub-clusters B, C and D) within the same 10 SNP cluster. Sub-cluster B (highlighted pink) – contaminated lamb mince; sub-cluster C – contaminated lamb sausage (highlighted yellow); sub-cluster D (highlighted blue) – direct contact with sheep or lambs or their environment in the northwest of England. Food isolates are highlighted with a red asterisk.
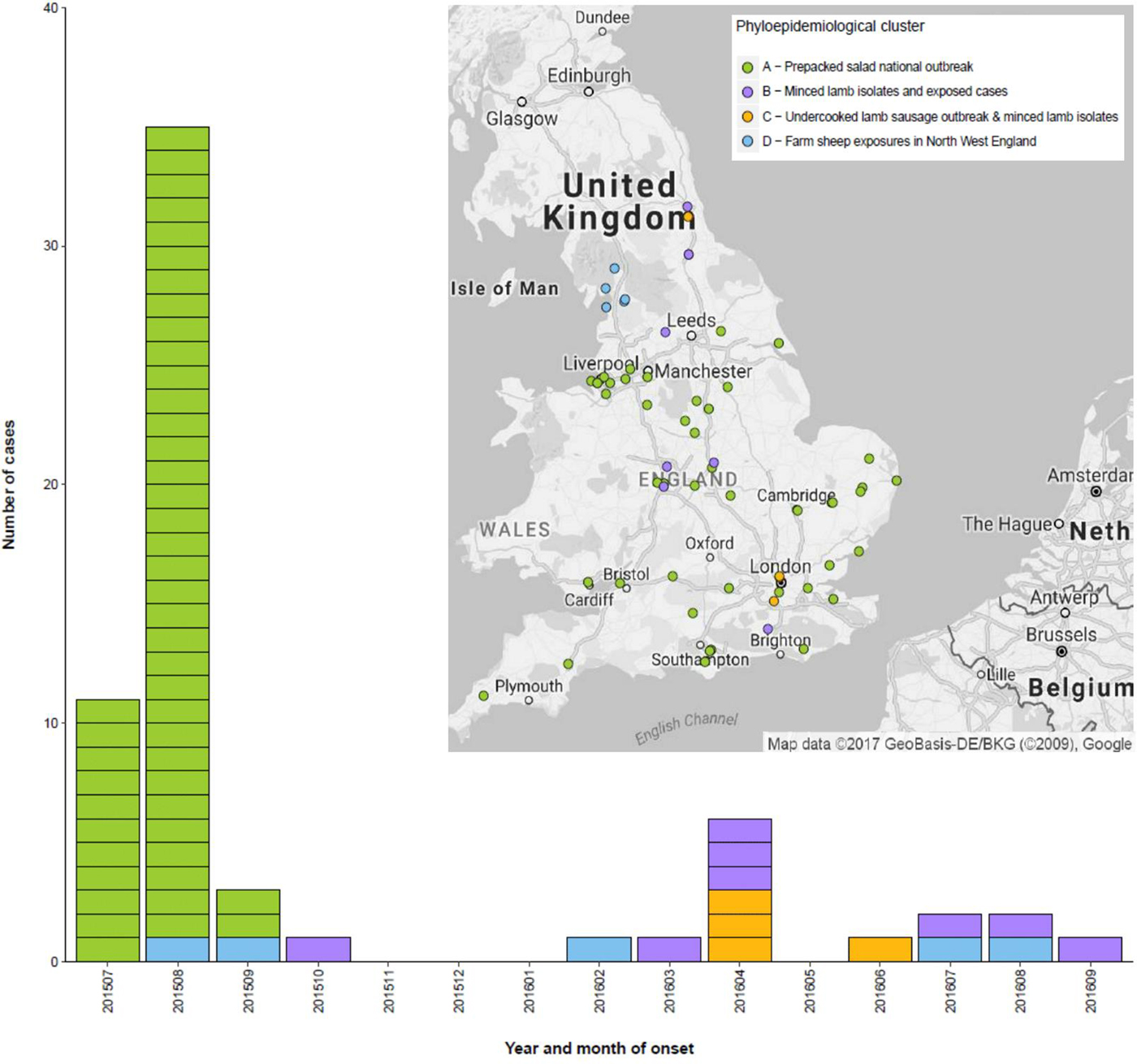
Fig. 2. Temporal and geographic distribution of cases within the 10 SNP cluster incorporating the outbreak attributed to consumption of contaminated prepacked salad and sporadic cases detected following the outbreak, UK, July 2015 – December 2016. Sub-cluster A (highlighted green) – salad outbreak; sub-cluster B (highlighted pink) – contaminated lamb mince; sub-cluster C – contaminated lamb sausage (highlighted yellow); sub-cluster D (highlighted blue) – direct contact with sheep or lambs or their environment in the northwest of England.
An exposure frequency analysis conducted on SESQs revealed that the most common exposure reported by cases was consumption of prepacked salad (38/47; 81%) and of these, 22 (58%) had purchased their prepacked salad at the same national supermarket chain (retailer X). The case–case study included 36 cases and 78 non-outbreak controls from the NESSS database. Results of the final multivariable model are shown in Table 1. Consumption of prepacked salad from retailer X was identified as the primary exposure associated with the outbreak (OR 54, 95% CI 11–247).
Table 1. Odds ratios for multivariable analysis of the outbreak associated with consumption of contaminated salad

* Number and proportion of primary cases exposed were calculated from a total of 45 cases. Of these, 36 were included in the case–case study and compared to 78 controls to derive the odds ratios displayed above.
Trace back and environmental investigations of the outbreak associated with consumption of contaminated salad
A focused trawl was then carried out on the first 24 cases to obtain further details on the type of prepacked salad consumed. Two salad product lines (bistro salad and mild mixed leaf salad) were implicated. A trace back investigation revealed that both salads shared an ingredient (green multi-leaf lettuce) and were processed and packed by a single distributor in the North West of England. The distributor was supplied by five UK salad growers (Fig. 3). The five source farms were visited and inspected. A review of routine test results of the produce revealed that high indicator E. coli levels had been detected in packaged produce from one farm in August 2015 during the outbreak. In addition, three farms used untreated irrigation water from open ponds (n = 2) or rivers (n = 1). Potential risks, or actual evidence, of animal ingress (wildlife) was identified at all farms. Follow-up of all potential areas of sub-optimal practice was undertaken by local or national enforcement authorities.

Fig. 3. Analysis of sheep movement data showed that lamb flocks sourced for the contaminated lamb meat products at farms C and D, and a network of lamb and sheep movement in the northwest of England, originated from a single 20 km radius area incorporating farms A and B. Green circles represent the location of the salad growers implicated in the outbreak sub-cluster A.
Microbiological examination of animal faecal specimens, food and environmental samples
Sampling and microbiological testing of the packaged products were undertaken from a selection of supermarkets visited by cases. In addition, lettuce leaves, chlorine wash water and environmental swabs from cutting and conveying equipment at the processing plant were tested for STEC. STEC was not detected in any of the samples tested, and no episodes of gastrointestinal illness had been reported from workers at the processing plant during the period of interest. A total of 32 salad leaf samples and 13 irrigation water samples from the source farms were tested, and STEC was not detected in any of the samples.
Phylogenetic relationships between isolates from outbreak and post-outbreak cases
After the outbreak was declared over, analysis of the WGS data identified 22 additional isolates that fell within a 10 SNP single linkage cluster of the outbreak strain. Of these, 17 isolates were from symptomatic human cases, and five isolates were from food products made from lamb meat sent for routine testing to a commercial food microbiology laboratory from a national retailer. Of the 17 human cases, three reported onset of symptoms between August and October 2015, and 14 reported onset of symptoms between March 2016 and September 2016 (Fig. 2).
Using a combination of epidemiological data and phylogenetic analysis, the outbreak and post-outbreak cases belonging to this 10 SNP single linkage cluster (SNP address: 18·35·397·765·1009.%) were categorised into four sub-clusters (Fig. 1). Sub-cluster A (highlighted in green in Figs 1 and 2) comprised 47 cases associated with the salad outbreak in 2015 (SNP variation 0–1). Sub-cluster B contained 11 isolates (SNP variation 0–4) received at the reference laboratory between October 2015 and September 2016. Sub-cluster B (highlighted in pink in Figs 1 and 2) included three isolates detected in lamb meat and eight geographically dispersed cases (Fig. 2), suggesting a nationally distributed food product was associated with infection. Seven of the eight cases reported consumption of sausages and/or burgers prior to onset of symptoms, of which four specifically reported consumption of food products made from lamb meat. Sub-clusters A and B were separated by a minimum distance of three SNPs (Fig. 1) and were estimated to share a common ancestor approximately 1·6 years ago, in early 2015 (95% HPD 1·5–2·9). Sub-cluster C (highlighted in yellow in Figs 1 and 2) comprised six isolates; four human cases linked to an outbreak caused by the consumption of undercooked lamb sausages at the same venue and two isolates from food products made from lamb meat (SNP variation 0–4). Sub-cluster D (highlighted in blue in Figs 1 and 2) was ancestral to the other sub-clusters and comprised five cases all resident in the northwest of England (SNP variation 8–25) (Fig. 2). The most-recent common ancestor of four sub-clades was estimated to the middle of 2014 (95% HPD 1·8–5·3).
Epidemiological investigation of the links between cases in the 10 SNP cluster
Links between the sub-clusters were investigated in order to identify a common epidemiological source and clarify transmission routes. A second case–case study comprising cases detected from 2016 onwards that were not part of the outbreak associated with the consumption of contaminated salad leaves, revealed that ovine exposures, including direct contact with sheep or lambs, and/or the handling and consumption of lamb meat (particularly minced meat products), were most strongly associated with illness (n = 16; aOR 8·24; 95% CI 1·55–39·74). Other exposures that met the criteria for inclusion in the final model (consumption of sandwiches, prepacked salad, cured meat and raw vegetables) were not significant (Table 2).
Table 2. Odds ratios for multivariable analysis of non-outbreak cases phylogenetically linked (10 SNP single linkage cluster) to an outbreak caused by consumption of contaminated salad to test an ovine exposure hypothesis

* Number and proportion of primary cases exposed were calculated from a total of 18 cases. Of these, 16 were included in the case–case study and compared to 1919 controls to derive the odds ratios displayed above.
Details of the movements of sheep and lambs to and from farms and other locations were supplied for a total of 30 sites associated with lambs being held prior to slaughter or cases having direct or indirect contact with lambs and/or sheep comprising a total of 1238 sheep in 89 independent movements between sites. The analysis revealed two distinct networks of sheep movements originating from two farms (farms A and B) within 20 km of each other on the Scottish–English border (Fig. 3). The network in the northwest of England developed from a single movement of 22 sheep from farm A in Scotland. The second network in the Welsh border region, developed from three independent movements of 138 sheep from farm B in Cumbria. Each movement contained 50, 38 and 50 animals, respectively.
Product trace back and analysis of sheep movement data showed that the lambs associated with the contaminated lamb sausage were sourced from farm C in the southeast of England. The lambs from farm C were directly linked to farm D on the Welsh–English border. Farm D was the source of the lambs associated with the contaminated mince (Fig. 3).
DISCUSSION
In England over the last 10 years, there has been an increase in reports of STEC O157:H7 outbreaks associated with raw vegetables or salad leaves [Reference Launders4–Reference Launders7]. Investigations of these outbreaks can be challenging, as they are often small in case numbers, geographically dispersed and the contaminated product is often a minor component of a meal that people may not remember. In addition, the source of the leaves used and the composition of bagged salad products changes frequently and distribution chains may be complex, making trace back to specific farms extremely difficult. Furthermore, salad produce has a short shelf-life and may contain very low levels of bacteria that are not detectable by microbiological testing [Reference Jenkins8]. Routine microbiological monitoring performed by the food business operator includes tests for indicator bacteria used to detect and estimate the level of faecal contamination, but are not specific for STEC O157:H7.
Increased awareness of consumption of contaminated salad leaves as a potential source of STEC O157:H7 infection has improved the quality of food history data from cases recorded on the SESQ in England. In this outbreak, pre-packed salad was quickly identified as the most likely source in the initial phase of the investigation, and this was confirmed by the case–case study. This was the first time that SESQs had been used for an analytical study. This approach reduces recall error, because questioning is completed relatively quickly after initial presentation, and reduces recall and interviewer bias because at the time of interview, the case would not have been classified as an outbreak case. However, using the SESQs for a case–case study does risk overmatching and does not allow the questions to be modified to precisely match the hypothesis, both of which may reduce study power to detect a real association. During this study, despite the risk of a high salad exposure in the control group, the outbreak strain cases were 54 times more likely to have consumed prepacked salad (95% CI 11–247). Detailed follow-up of the cases confirmed the hypothesis and directed the trace back investigation to focus on green multi-leaf lettuce packed by a single supplier who received produce from five farms.
This investigation was the first time that data from ARAMS has been used to investigate epidemiological links between phylogenetically related isolates of STEC O157:H7. The analysis was limited because at the time of this investigation the ARAMS did not hold a complete record of all sheep movements that had occurred during the time frame of interest. In particular, data were not available for the sheep holdings in the vicinity of the growers irrigating salad crops with untreated water. Ultimately, a definitive link could not be established between sub-cluster A (cases associated with the consumption of contaminated salad leaves) and the other three sub-clusters. However, a direct link was established between the isolates in sub-clusters B (cases associated with the consumption of contaminated lamb mince) and C (cases associated with the consumption of contaminated lamb sausage) via the movements of lambs between farm C in the southeast of England and farm D on the Welsh borders. A potential geographical link was identified between sub-clusters B (cases associated with the consumption of contaminated lamb mince) and D (cases associated with direct contact with lambs and/or sheep and their environment) as their associated sheep movement networks could be traced back to two farms 20 km apart.
Animal faeces may be distributed over wide areas by stream and river systems following heavy rainfall. More timely and intensive sampling of the location of the farms where the green multi-leaf lettuce was produced, including the irrigation water and cattle and/or sheep grazing upstream of the irrigation point, may have facilitated the detection of the outbreak strain in these animals, or in their environment [Reference Söderström21]. In a previous outbreak associated with consumption of contaminated watercress, although not detected in the watercress, STEC O157:H7 was successfully detected in water used to irrigate the crop on the farm [Reference Jenkins8]. During an investigation of an outbreak of STEC O157:H7 caused by contaminated raw drinking milk, the causative organism was not detected in the milk but was isolated from faecal pats from the cows producing the milk [Reference Butcher9].
Microbiological testing in response to the outbreak failed to identify the pathogen in food collected during the investigation. More appropriate testing using PCR assays targeting the stx genes may enable the identification of contaminated of ready-to-eat vegetables, including salad vegetables, before and after they enter the food chain. Salad vegetables are considered ready to eat food products. Washing during and after processing is largely ineffective in removing bacteria, particularly those that are internalised, the most important food safety controls are those applied in the field [Reference Shaw22]. In England, salad production is highly localised to small parts of the country, and some areas are geographically close to areas of high animal density, particularly in the northwest and southwest of the country. The likelihood of direct or indirect contamination of ready to eat salad crops (e.g. from birds, wildlife or contaminated watercourses) from the animal reservoir is likely to be higher in these areas. Farms growing salad vegetables are deemed primary producers. The legal requirements for hygiene in primary production are laid out in EC Regulation 852/2004 which applies the principles of good agricultural practice and good hygiene practice but are not as stringent as those required for food processing and production. However, business must be able to demonstrate that their operations are managed in a way that control food safety risks, including those associated with the use of water, and primary growers must use potable or clean water to prevent contamination.
During this study, the potential for identifying the geographical origin of an outbreak strain by analysing epidemiological data associated with cases in phylogenetically related sub-clusters was explored. Although, a robust link between the outbreak sub-cluster (A) and the other three ovine associated sub-clusters B, C and D (linked to consumption of lamb mince, consumption of lamb sausage and direct contact with sheep or lambs, or their environment, respectively) was not confirmed, epidemiological links were established between three ovine sub-clusters by mapping the recent movement of sheep and lambs across the UK. Given the close phylogenetic relationship between the outbreak strain and the isolates from cases with ovine exposures, it is plausible that ovine faeces may have contaminated the salad leaves via untreated irrigation water or run-off from fields nearby. Timely and targeted veterinary and environmental sampling should be considered during outbreaks of STEC, particularly where ready to eat vegetables are implicated.
SUPPLEMENTARY MATERIAL
The supplementary material for this article can be found at https://doi.org/10.1017/S0950268817002874.
ACKNOWLEDGEMENTS
The authors would like to thank Neil Perry, Vivienne do Nascimento and Marie Chattaway at GBRU, Kate James from PHE WM Centre and colleagues at PHE Food, Water and Environmental Microbiology Laboratories and Preston, Porton and Birmingham. The authors would also like to acknowledge everyone who was part of the Outbreak Control Team including Drazenka Tubin-Delic, Jonathan Lighthill and Joanne Edge the Food Standards Agency, Paul Homes, Alan Wright and Charlotte Featherstone at the Animal and Plant Health Agency and Eamon Rodgers at Worcestershire Regulatory Services County Council. This work was supported by the National Institute for Health Research Health Protection Research Unit in Gastrointestinal Infections. Jeremy Hawker, Richard Elson, Paul Cleary, Tim Dallman and Tom Inns are affiliated to the National Institute for Health Research Health Protection Research Unit (NIHR HPRU) in Gastrointestinal Infections at University of Liverpool in partnership with PHE, in collaboration with University of East Anglia, University of Oxford and the Institute of Food Research. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR, the Department of Health or PHE.








