Introduction
On 14 January 2013, the US Food and Drug Administration (FDA) announced permission for a multiplex nucleic acid test, the xTAG® Gastrointestinal Pathogen Panel (GPP) (Luminex Corporation, USA), which simultaneously detects 11 common viral, bacterial and parasitic causes of infectious gastroenteritis, to be marketed in the USA. This announcement reflects the current move towards the development and commercialization of detection technologies for the diagnosis of infectious syndromes whether, enteric [Reference Claas1–Reference Stark6], respiratory [Reference Anderson7–Reference Sakthivel13], sexually transmitted [Reference Lee14], affecting the central nervous system [Reference Shin15] or causing sepsis [Reference Blaschke16, Reference Josefson17]. Commercially available tests for the detection of specific pathogens associated with urinary tract infection, neonatal infection or infection in the immunocompromised host have also been developed by commercial diagnostic companies including Fast-track Diagnostics (Junglinster, Luxembourg), AusDiagnostics Pty Ltd (Sydney, Australia) and Seegene (Seoul, Korea).
The superior sensitivity, specificity, and reproducibility of nucleic acid amplification techniques combined with the ability to identify a broader range of human pathogens in a rapid format has driven the move from classical diagnostic microbiological techniques including microscopy, microbial culture, antigen detection and serology within the diagnostic setting [Reference Amar18–Reference Weinberg23]. Moreover, the ability to reduce the myriad of techniques utilized within the routine diagnostic setting such as culture, including selective media and enrichment, biochemical identification, microscopy including immunofluorescence, antigen detection by enzyme immunoassay or particle agglutination as well as the delayed serological diagnosis of infectious disease, can streamline sample throughput. Commercial molecular diagnostic technologies encompassing nucleic acid extraction, liquid handling, molecular amplification and identification, which are amenable to partial or complete automation and high throughput, have become increasingly available [24]. However, it is important to understand the limitations of any new technology as well as the clinical relevance of the results obtained. Microbial culture and microscopy can be catch-all methods for detection of a broad spectrum of pathogens including new or unanticipated agents with culture identifying and isolating viable organisms for further study [Reference Leland and Ginocchio25, Reference Ogilvie26]. Nucleic acid amplification technologies, which are limited to existing knowledge of a microorganism's genome, are highly specific and do not discriminate between viable and dead organisms with resultant detection of microbes that are present at non-pathogenic levels. Thus, care is required when designing a molecular amplification assay and interpreting its results [Reference Mackay27]. Furthermore, it is common for microbial genomes to contain unexpected mutations, which may render a molecular assay ineffective [Reference Mackay27], whereas microbial culture, antigen detection and serology are less likely to be influenced by mutations unless they result in phenotypic changes.
Detection technologies for the diagnosis of syndromic infection range from a number of grouped monoplex real-time polymerase chain reaction (PCR) and real-time reverse-transcription (RT)–PCR assays allowing for the detection of individual pathogens to multiplex real-time PCR and RT–PCR assays with the capacity to detect up to six analytes simultaneously using the same thermal cycling profile. However, the low multiplexing capability of real-time PCR instrumentation represents a major drawback of the technology within the clinical setting [Reference Reddington28], which is limited to four or five fluorescent channels detecting fluorescent dyes at different wavelengths or using melting-point analysis to differentiate among PCR products. Limitations are associated with the number of detectors available or the need for discrimination in detectable wavelengths to prevent cross-talk.
The increasing availability of commercial multiplex nucleic acid detection tests for the diagnosis of infectious syndromes
Recent advances within the clinical diagnostics market has seen an increase in the availability of Conformité Européene – in-vitro diagnostic (CE-IVD)-approved, multiplex real-time PCR and RT–PCR kits for the simultaneous detection of multiple targets in the same reaction vessel using commonly available real-time PCR platforms especially for respiratory viruses (Table 1). However, a shift towards the diagnosis of infectious syndromes has driven commercial diagnostic companies to develop multiparametric molecular diagnostic tests within other disease categories. Indeed, Fast-track Diagnostics (Luxembourg) offer optimized multiplexed real-time PCR and RT–PCR primer and probe mixes based on wide-ranging infection syndromes using the same thermal cycling profile. AusDiagnostics Pty Ltd (Australia) also offer highly multiplexed panels using multiplex tandem PCR, which employs two sequential steps. Step 1 is a short (15 cycles) multiplexed pre-amplification reaction using primers homologous to all targets in the panel so competition between primers is avoided allowing low concentrations of targets to be detected, even when multiple pathogens are present. Reverse transcriptase is included in the step 1 reaction for panels including RNA targets. A prior cDNA synthesis reaction is not necessary. In step 2, the product from step 1 is diluted into individual wells for real-time PCR reactions using primers ‘nested inside’ the primers used for step 1. The Easy-Plex™ liquid handling robot (AusDiagnostics, Pty Ltd) automates this process. The step 2 PCR reaction is performed in the Rotor-Gene 6000 real-time analyser. DNA amplification is measured by the increase in fluorescence when Eva-Green™ dye (Biotium Inc., USA) is incorporated into the DNA being formed in the specific amplification reaction.
Table 1. Examples of commercially available multiparametric detection technologies for diagnosis of respiratory infection
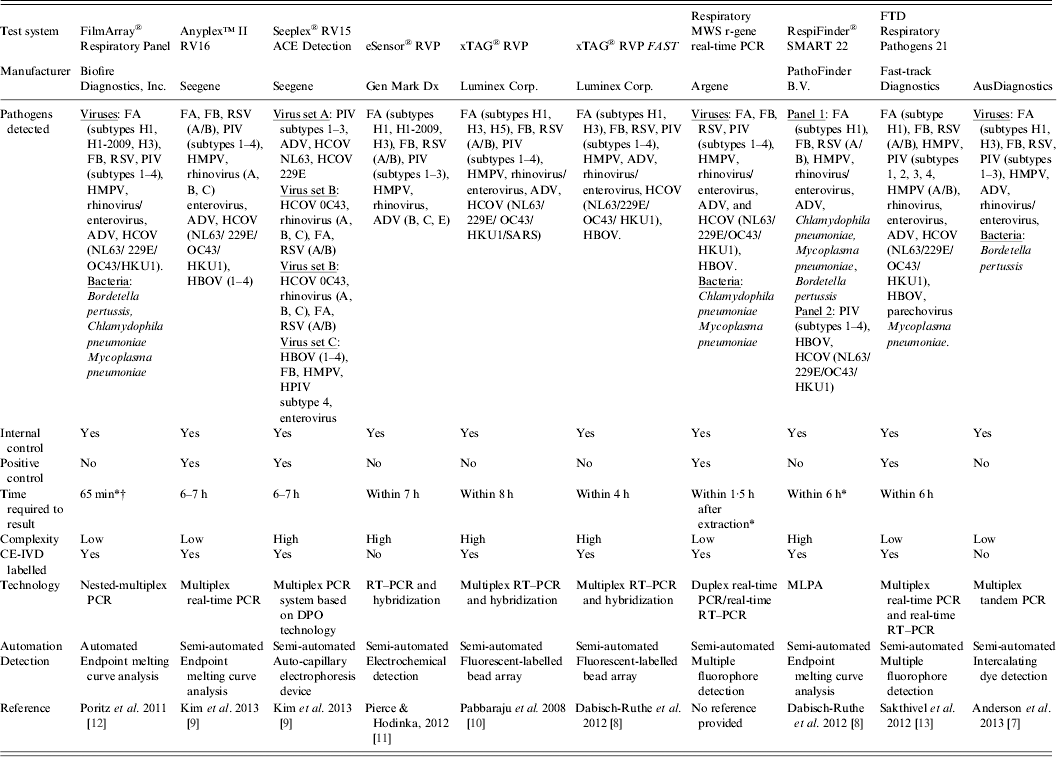
ADV, Adenovirus; CE-IVD, Conformité Européene – in-vitro diagnostic; DPO, dual-priming oligonucleotides; FA, influenza virus type A; FB, influenza virus type B; HBOV, human bocavirus; HCOV, human coronavirus; HMPV, human metapneumovirus; MLPA, multiplex ligation-dependent probe amplification; MWS, multi-well system; PCR, polymerase chain reaction; PIV, parainfluenza virus; RSV, human respiratory syncytial virus; RT–PCR, reverse transcription–polymerase chain reaction; RVP, respiratory virus panel.
* Time required to obtain result was taken from commercial diagnostic company product literature.
† The FilmArray integrates sample preparation, amplification, detection, and analysis into one simple system that requires 2 min of hands-on time and has a total run time of about 1 h.
Seegene (Korea) utilize Dual-Priming Oligonucleotides (DPO™) in the Seeplex® multiplex PCR and Anyplex™ multiplex real-time PCR product ranges in combination with detection by automatic capilliary electrophoresis or melting-curve analysis utilizing Tagging Oligonucleotide Cleavage and Extension (TOCE™) technology, respectively, which allow high multiplexing by enabling ‘one channel, many targets'. These methods are highly appropriate for the identification and differentiation of viral and bacterial pathogens with very variable genetic characteristics and low availability of primer sites [Reference Shin15].
The benefits that multiplexed nucleic acid detection tests provide to the diagnosis of infection syndromes over conventional techniques within the clinical microbiology setting are increasingly highlighted, especially in relation to gastroenteritis [Reference Claas1–Reference Stark6, Reference Mengelle29, Reference Navidad30]. The enteric targets included in multiplex molecular diagnostic tests available on the clinical diagnostics market vary considerably (Table 2). The CE-IVD marked EntericBio® Real-Time Gastro Panel I (Serosep Ltd, Ireland) currently only targets Salmonella enterica spp., Shigella spp., Campylobacter spp., and Shiga toxin-producing Escherichia coli (STEC) (Table 2). However, unique to EntericBio Real-Time Gastro Panel I is the ability to perform the test directly from the stool sample thereby removing the usual requirement for the nucleic acid purification, which often presents a bottleneck to the successful implementation of nucleic acid detection within a routine virological setting [Reference Niesters31]. The entire master mix required to perform each test is lyophilized into individual reaction wells, which offers an additional improvement to the workflow within the routine diagnostic setting. In combination, the exceptional features of the EntericBio Real-Time Gastro Panel I enable results to be generated within 3 h [Reference Koziel4] (Table 2). In contrast, the CE-IVD marked Seeplex® Diarrhea ACE Detection multiplex PCR system based on DPO technology (Seegene, Korea) can detect four viruses and/or 10 bacteria using a virus panel (panel V) and two bacterial panels, bacterial panel 1 (panel B1) and bacterial panel 2 (panel B2) [Reference Coupland3] (Table 2). There are several limitations associated with the Seeplex Diarrhea ACE Detection system. First, no option is available within the current system for the detection of human diarrhoeal parasites and although the Seeplex system incorporates quality controls, the internal control is only available for inclusion in each PCR master mix, which does not allow validation of the nucleic acid extraction or reverse transcription processes. Moreover, reverse transcription is performed as a separate step, which in turn increases the duration of the assay [Reference Coupland3]. The average turnaround time to process 96 samples using the Seeplex system was 9–10 h or 0·6 h per target in a run of 96 samples compared to 24–48 h [Reference Coupland3].
Table 2. Examples of commercially available multiparametric detection technologies for diagnosis of infectious gastroenteritis
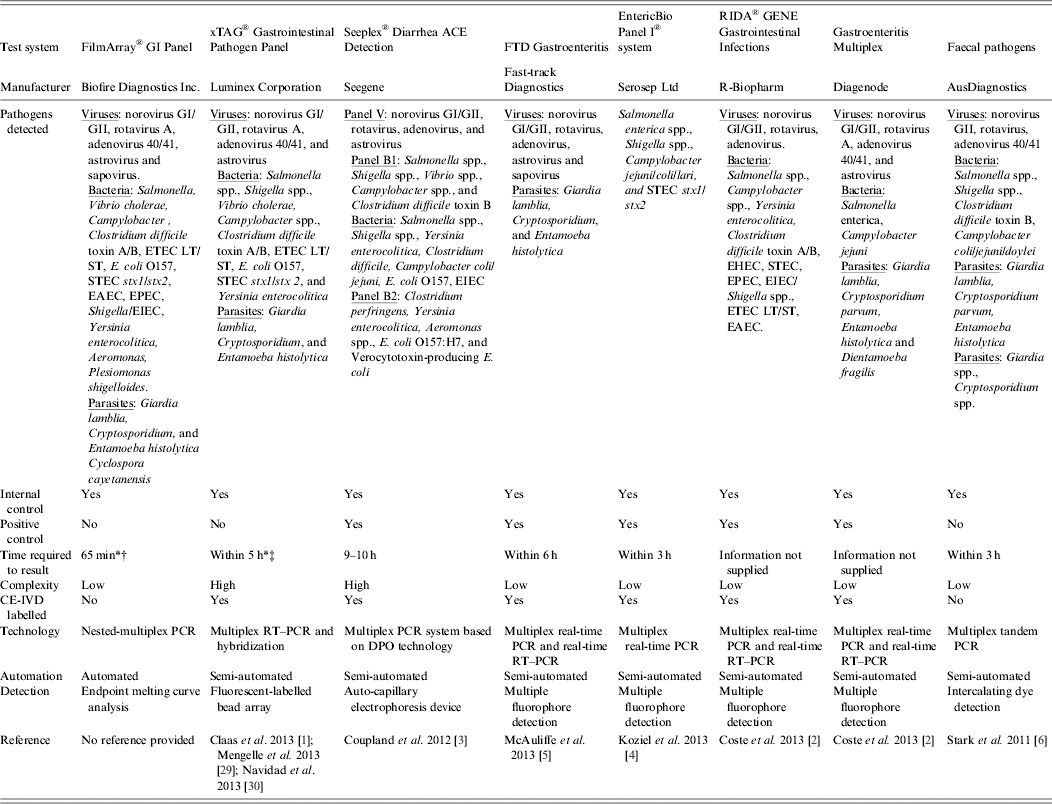
DPO, Dual-priming oligonucleotides; EAEC, enteroaggregative Escherichia coli; EHEC, enterohaemorrhagic E. coli; EIEC, enteroinvasive E. coli; EPEC, enteropathogenic E. coli; ETEC LT/ST, enterotoxigenic E. coli LT/ST; PCR, polymerase chain reaction; RT–PCR, reverse transcription–polymerase chain reaction; STEC stx1/stx2, Shiga-like toxin-producing E. coli stx1/stx2;
* Time required to obtain result was taken from commercial diagnostic company product literature.
† The FilmArray integrates sample preparation, amplification, detection, and analysis into one simple system that requires 2 min of hands-on time and has a total run time of about 1 h.
‡ Time estimate of within 5 h is for a maximum of one extraction (24 samples), unless multiple extractors are available. Does not include pre-treatment.
The CE-IVD marked xTAG GPP (Luminex Corporation, USA), and FilmArray® GI Panel (BioFire Diagnostics Inc., USA), which is currently available for research use only, provide the most comprehensive commercial multiplex molecular diagnostic tests available for gastroenteritis diagnosis. The xTAG GPP can simultaneously detect and identify three viruses, nine bacteria and three parasites while the FilmArray GI Panel tests for a panel of five viruses, 14 bacteria and four parasites (Table 2). However, there are significant differences between these tests. The xTAG GPP procedure incorporates sample pre-treatment, which consists of bead beating in lysis buffer to ensure maximum efficiency during nucleic acid purification, multiplex RT–PCR, bead hybridization and detection using the proprietary universal tag sorting system and data acquisition and analysis. The workflow can be completed within 5 h, which is based on one extraction of 24 samples but does not include the pre-treatment stage. In contrast, the FilmArray system brings sample to results in about 1 h, with minimal hand-on time to process. However, a significant drawback of the FilmArray system is its low throughput, as only a single sample can be processed on the instrument at one time, which limits the overall utility of the test in laboratories with moderate to high numbers of specimens to be tested [Reference Pierce32].
There are also an increasing number of multiparametric molecular diagnostic tests available for the diagnosis of meningitis and sexually transmitted disease (STD). The Seeplex® Meningitis ACE Detection (Seegene, Korea) multiplex PCR system based on DPO technology detects 12 common bacterial and viral causes of acute meningitis, i.e. Streptococcus pneumoniae, Haemophilus influenzae, Neisseria meningitidis, group B streptococcus, Listeria monocytogenes, herpes simplex virus types 1 and 2, cytomegalovirus, Epstein–Barr virus, varicella zoster virus, human herpes virus 6 and human enterovirus [Reference Shin15]. DPO technology is also utilized in the Seeplex® STD6 ACE Detection system, which is designed to simultaneously detect six STD pathogens, i.e. Trichomonas vaginalis, Mycoplasma hominis, Mycoplasma genitalium, Ureaplasma urealyticum, Chlamydia trachomatis, Neisseria gonorrhoeae [Reference Lee14]. Similarly, the Anyplex™ II STI-7 Detection system detects the aforementioned STD pathogens plus Ureaplasma parvum using DPO and TOCE technology in a single real-time PCR.
The advantages and challenges of multiplex nucleic acid detection tests
Despite the differences between these multiparametric molecular diagnostic tests all have in common the ability to provide a more comprehensive assessment of the aetiology of disease [Reference Claas1] due to increased diagnostic yield compared to conventional diagnostic tests [Reference Oosterheert33, Reference van de Pol34]. These tests can also accelerate the microbial detection/identification phase of the laboratory diagnostic cycle to meet the critical 6- to 24-h window [Reference Bissonnette and Bergeron35]. The ability to rapidly detect and distinguish multiple potentially infectious pathogens is critical for the accurate diagnosis of seasonal and sporadic outbreaks, emerging pathogens and agents of bioterrorism [Reference Poritz12]. Clinical syndromes are seldom specific to a single pathogen, so detection strategies that allow multiple agents to be simultaneously considered [Reference Briese36] can have a significant impact on infectious disease management since multiparametric molecular diagnostic tests can provide a more accurate representation of the true pathogen spectrum in clinical samples [Reference Brunstein37].
In the absence of rapid tests, infections are managed using empirical antibiotic regimens, which are associated with overuse of broad-spectrum antibiotics, which has major implications for the development of bacterial resistance and emergence of hospital-acquired infections [Reference Bissonnette and Bergeron35]. Inadequate or inappropriate antimicrobial treatment and the delayed administration of appropriate antimicrobial therapy correlate with negative clinical outcomes in patients with bacteraemia and sepsis compared to patients who receive appropriate therapy from the onset [Reference Lueangarun and Leelarasamee38]. This was reflected in a longer hospital stay, a higher risk of Clostridium difficile-associated infection, excess mortality and higher cost of therapy per bacteraemic episode [Reference Lueangarun and Leelarasamee38–Reference Shehab45].
Rapid diagnosis can have a major impact on patient care and outcome, most importantly significant reductions in hospital stay, inappropriate or unnecessary antibiotic use, and associated common adverse reactions including rash, abdominal pain, diarrhoea and vomiting as well as informing decisions regarding infection control measures [Reference Barenfanger, Drake and Kacich46–Reference Woo52]. However, most studies have concentrated on the diagnostic capabilities of multiparametric molecular diagnostic tests [Reference McAuliffe5, Reference Chen53, Reference Tsalik54] while the clinical and economic impact of these tools has received limited attention [Reference Oosterheert33]. Oosterheert et al. [Reference Oosterheert33] conducted a randomized controlled trial involving 107 adults with lower respiratory tract infections at two hospitals in The Netherlands. All patients had upper respiratory tract specimens tested for viral and atypical bacterial pathogens by real-time PCR as well as by conventional diagnostic procedures, but only results for patients in the intervention group were reported to the treating physician; results for patients in the control group were unavailable. The implementation of multiparametric detection technologies for diagnosing respiratory infections increased the diagnostic yield compared to conventional diagnostic tests but did not reduce antibiotic use, antibiotic costs, or the duration of hospital stay [Reference Oosterheert33]. Wishaupt et al. [Reference Wishaupt55] also found that RT–PCR testing had a high yield of viral diagnoses, but rapid communication did not lead to decreases in hospital admissions, shorter hospital stays, or less antibiotic use for children with acute respiratory infections. In contrast, Brittain-Long et al. [Reference Brittain-Long56] demonstrated that access to a rapid molecular diagnostic tool for aetiological diagnosis of viral respiratory infection significantly reduced antibiotic prescriptions at the initial visit in a primary-care setting but this effect was no longer evident at follow-up. These studies highlight the difficulties in evaluating the impact of molecular diagnostic tests on patient management, especially in relation to respiratory tract infection since bacterial co-infection is associated with about 40% of viral respiratory tract infections requiring hospitalization [Reference Falsey57]. Hence, clinicians are unwilling to alter therapy based on discovery of a viral pathogen [Reference Murdoch58]. Fortunately, new rapid multiplex molecular diagnostic tools are becoming available, which are designed to detect bacterial respiratory pathogens. These include the Anyplex™ II RB5 Detection system (Seegene, Korea), which detects and differentiates the most common causes of atypical pathogens, Mycoplasma pneumoniae, Chlamydophila pneumoniae, Legionella pneumophila and two causative agents of whooping cough, the commonly detected Bordetella pertussis as well as the less common but vaccination-ineffective Bordetella parapertussis. Fast-track Diagnostics (Luxembourg), Pathofinder B.V. (The Netherlands) and AusDiagnostics Pty Ltd (Australia) also offer multiplex PCR assays for the simultaneous detection and differentiation of relevant respiratory viruses and bacteria associated with respiratory tract infection.
The LightCycler® SeptiFast Test MGRADE (Roche Diagnostics, Switzerland), a commercial real-time PCR designed to detect and identify 25 bacterial and fungal species that comprise >90% of the pathogens causing bloodstream infections in critical care [Reference Dark, Dean and Warhurst59] epitomizes the difficulties associated with changing the clinical management of infectious disease. The LightCycler SeptiFast MGRADE test was the first PCR-based system to be awarded a CE mark for pathogen detection and identification in blood samples and, to date, is the most intensively investigated multiplex real-time PCR assay in the clinical setting of sepsis [Reference Dark, Dean and Warhurst59]. It offers demonstrable diagnostic value in terms of enhanced detection of the most common pathogen species in patients with suspected sepsis and for the timely diagnosis of bloodstream infections, particularly in antibiotic pre-treated patients [Reference Westh60, Reference Yanagihara61]. The LightCycler SeptiFast Test MGRADE system is now part of a clinical diagnostic validation study to determine whether this multiplex molecular diagnostic technology has sufficient clinical diagnostic accuracy, which represents a crucial phase of detailed independent health technology assessment of the first multiplex real-time PCR technique aimed at helping deliver more effective care to critically ill patients internationally [Reference Dark62].
The extraordinary sensitivity associated with molecular diagnostic tests also brings a new set of challenges that include detection of dead microbes or potential pathogens that simply colonize non-sterile sites [63]. Certainly, quantitative or semi-quantitative molecular methods are utilized to establish a clinically significant result for viral infection as asymptomatic infection is associated with a significantly lower viral load [Reference Gerna64, Reference Phillips65] and may help to separate bacterial colonization from disease [63]. It seems that only with this information can clinicians make well-informed decisions, which would promote judicious antibiotic use and permit pathogen-targeted antibacterial therapy [63]. Multiparametric diagnostic tests may also be augmented by the inflammatory biomarker procalcitonin (PCT) since a growing body of evidence supports PCT use to differentiate bacterial from respiratory viral diagnoses, which may improve individualized decision-making regarding antibiotic treatment when multiparametric molecular diagnostic tests cannot exclude the possibility of bacterial superinfection [Reference Schuetz, Raad and Amin66].
Multiparametric detection technologies can come with a price in human resources and qualified technical staff [Reference Bissonnette and Bergeron35]. Nevertheless, multiplex nucleic acid detection tests also offer the opportunity to adapt clinical microbiology services within the current austere environment through rationalizing or redistributing labour and costs while maintaining and improving the provision of routine diagnostic services [Reference Dundas67]. Common to all multiplex nucleic acid detection tests and recently demonstrated using the Luminex xTAG respiratory virus panel is the ability to increase laboratory efficiency by reducing hands-on time and operational steps while standardizing workflow in comparison to viral direct immunofluorescence assay (DFA) and culture [Reference Dundas67]. The cost–benefit studies constructed using multiplex PCR demonstrate savings in the absence of reduced rates of antibiotic usage. Multiplex PCR testing for respiratory viruses achieved using the xTAG respiratory virus panel test was the least costly strategy for the diagnosis of respiratory virus infections compared to standard non-molecular diagnostic methods such as viral culture and DFA [Reference Mahony68].
Conclusions
PCR-based technologies have become standard within the clinical laboratory setting over the last two decades. Experience of the clinical significance of results generated using these powerful molecular diagnostic tools has accumulated during this time and there are many benefits to be gained by utilizing sensitive, specific, and rapid molecular diagnostic techniques. However, the clinical utility of these techniques urgently needs to be determined through well-structured clinical trials comparing new with traditional methods. Diagnosis of syndromic infections represents a new pathway for the diagnosis of infection and newer molecular diagnostic tools, which will streamline workflows in the routine diagnostic setting, must incorporate all relevant pathogens in order to improve patient management.
DECLARATION OF INTEREST
None.



