I. INTRODUCTION
Vismodegib (sold under the brand name Erivedge) is used to treat unresectable or metastatic basal cell carcinoma. It belongs to a class of medicines referred to as hedgehog pathway inhibitors due to its ability to block the action of a protein that signals cancer cells to multiply (MedLine, 2022). The systematic name (CAS Registry Number 879085-55-9) is 2-chloro-N-(4-chloro-3-pyridin-2-ylphenyl)-4-methylsulfonylbenzamide. A two-dimensional molecular diagram is shown in Figure 1.
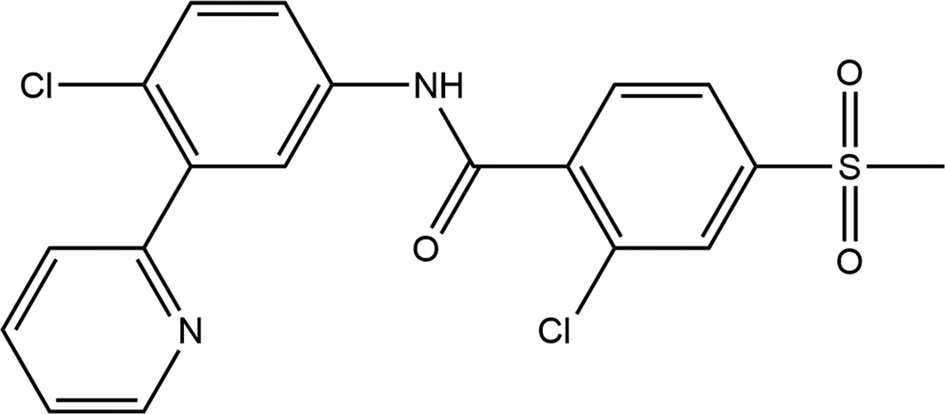
Figure 1. The 2D molecular structure of vismodegib.
Vismodegib and processes for its preparation were disclosed in US Patent 7,888,364 B2 (Gunzner et al., Reference Gunzner, Sutherlin, Stanley, Bao, Castanedo, Lalonde, Wang, Reynolds, Savage, Malesky and Dina2011; Curis, Inc.). The European Medicines Agency (EMA CHMP, 2013) reports that vismodegib exists primarily in the thermodynamically stable polymorph B. Polymorph A was discovered as the initial crystalline form, but could not be reproduced. Polymorph B is the only one which has been used in clinical development. A single-crystal structure was mentioned, but we found none in the open literature. Forms C and E were observed infrequently during extensive polymorph screening. Parthasaradhi Reddy et al. (Reference Reddy, K, Reddy, Muralidhara Reddy, Srinivas Reddy and Vamsi Krishna2014; Hetero Research Foundation) claimed a new polymorph of vismodegib (Form II) and provided powder diffraction data for Form I, which is described as the prior art of the ‘364 patent. We, therefore, presume that the Hetero Form I corresponds to EMA Form B. The mechanisms of action of vismodegib and clinical trials have been reviewed by Ruiz-Salas et al. (Reference Ruiz-Salas, Alegre, López-Ferrer and López-Ferrer2013). This work was carried out as part of a project (Kaduk et al., Reference Kaduk, Crowder, Zhong, Fawcett and Suchomel2014) to determine the crystal structures of large-volume commercial pharmaceuticals, and include high-quality powder diffraction data for them in the Powder Diffraction File (Gates-Rector and Blanton, Reference Gates-Rector and Blanton2019).
II. EXPERIMENTAL
Vismodegib was a commercial reagent, purchased from TargetMol (Lot #120202), and was used as-received. The white powder was packed into a 1.5 mm diameter Kapton capillary, and rotated during the measurement at ~50 Hz. The powder pattern was measured at 295 K at beamline 11-BM (Antao et al., Reference Antao, Hassan, Wang, Lee and Toby2008; Lee et al., Reference Lee, Shu, Ramanathan, Preissner, Wang, Beno, Von Dreele, Ribaud, Kurtz, Antao, Jiao and Toby2008; Wang et al., Reference Wang, Toby, Lee, Ribaud, Antao, Kurtz, Ramanathan, Von Dreele and Beno2008) of the Advanced Photon Source at Argonne National Laboratory using a wavelength of 0.458968(2) Å from 0.5 to 50° 2θ with a step size of 0.001° and a counting time of 0.1 s per step. The high-resolution powder diffraction data were collected using twelve silicon crystal analyzers that allow for high angular resolution, high precision, and accurate peak positions. A mixture of silicon (NIST SRM 640c) and alumina (NIST SRM 676a; Cline et al., Reference Cline, Von Dreele, Winburn, Stephens and Filliben2011) standard (ratio Al2O3:Si = 2:1 by weight) was used to calibrate the instrument and refine the monochromatic wavelength used in the experiment.
The pattern was indexed using N-TREOR (Altomare et al., Reference Altomare, Cuocci, Giacovazzo, Moliterni, Rizzi, Corriero and Falcicchio2013) on a primitive monoclinic unit cell with a = 16.8934, b = 10.1866, c = 12.1444 Å, β = 106.68°, V = 1979.79 Å3, and Z = 4. A reduced cell search in the Cambridge Structural Database (Groom et al., Reference Groom, Bruno, Lightfoot and Ward2016) yielded 18 hits, but no structures of vismodegib derivatives. The suggested space group was P21/a, which was confirmed by successful solution and refinement of the structure. The structure was solved by direct methods using EXPO2014 (Altomare et al., Reference Altomare, Cuocci, Giacovazzo, Moliterni, Rizzi, Corriero and Falcicchio2013). Some atom types had to be reassigned manually, and the hydrogen atoms were added using Materials Studio (Dassault, 2021).
Rietveld refinement was carried out using GSAS-II (Toby and Von Dreele, Reference Toby and Von Dreele2013). Only the 2.0–25.0° portion of the pattern was included in the refinement (d min = 0.886 Å). All non-H bond distances and angles (plus the plane of the fused ring system) were subjected to restraints, based on a Mercury/Mogul Geometry Check (Bruno et al., Reference Bruno, Cole, Kessler, Luo, Motherwell, Purkis, Smith, Taylor, Cooper, Harris and Orpen2004; Sykes et al., Reference Sykes, McCabe, Allen, Battle, Bruno and Wood2011). The Mogul average and standard deviation for each quantity were used as the restraint parameters. The restraints contributed 2.1% to the final χ 2. The hydrogen atoms were included in calculated positions, which were recalculated during the refinement using Materials Studio (Dassault, 2021). The U iso of the heavy atoms were grouped by chemical similarity. The two Cl atoms were refined anisotropically. The U iso for the H atoms were fixed at 1.3× the U iso of the heavy atoms to which they are attached. The peak profiles were described using the generalized microstrain model. The background was modeled using a 3-term shifted Chebyshev polynomial, and a peak at 6.65° 2θ was used to model the scattering from the Kapton capillary and any amorphous component.
The final refinement of 122 variables using 28 045 observations and 71 restraints yielded the residuals R wp = 0.1300 and GOF = 3.27. The largest peak (2.20 Å from C15) and hole (1.94 Å from C19) in the difference Fourier map were 0.50(12) and −0.53(12) eÅ−3, respectively. The largest errors in the difference plot (Figure 2) are in the shapes of many of the strong low-angle peaks, and suggest that the sample may have suffered beam damage during the measurement.

Figure 2. The Rietveld plot for the refinement of vismodegib. The blue crosses represent the observed data points, and the green line is the calculated pattern. The cyan curve is the normalized error plot. The red curve indicates the background. The vertical scale has been multiplied by a factor of 20× for 2θ > 13.0°. The row of blue tick marks indicates the calculated reflection positions.
The crystal structure was optimized using VASP (Kresse and Furthmüller, Reference Kresse and Furthmüller1996) (fixed experimental unit cell) through the MedeA graphical interface (Materials Design, 2016). The calculation was carried out on 16 2.4 GHz processors (each with 4 GB RAM) of a 64-processor HP Proliant DL580 Generation 7 Linux cluster at North Central College. The calculation used the GGA-PBE functional, a plane wave cutoff energy of 400.0 eV, and a k-point spacing of 0.5 Å−1 leading to a 2 × 2 × 2 mesh, and took ~64.7 h. A single-point density functional calculation (fixed experimental cell) and population analysis were carried out using CRYSTAL17 (Dovesi et al., Reference Dovesi, Erba, Orlando, Zicovich-Wilson, Civalleri, Maschio, Rerat, Casassa, Baima, Salustro and Kirtman2018). The basis sets for the H, C, N, and O atoms in the calculation were those of Gatti et al. (Reference Gatti, Saunders and Roetti1994), and those for S and Cl were those of Peintinger et al. (Reference Peintinger, Vilela Oliveira and Bredow2013). The calculations were run on a 3.5 GHz PC using 8 k-points and the B3LYP functional, and took ~2.6 h.
III. RESULTS AND DISCUSSION
The synchrotron powder pattern of this study matches the pattern for Form I reported by Parthasaradhi Reddy et al. (Reference Reddy, K, Reddy, Muralidhara Reddy, Srinivas Reddy and Vamsi Krishna2014) well enough to conclude that they represent the same material (Figure 3), corresponding to the thermodynamically stable Form B of vismodegib. The root-mean-square (rms) Cartesian displacement between the non-H atoms in the Rietveld-refined and DFT-optimized structures is only 0.055 Å (Figure 4), and the maximum difference is 0.118 Å. The excellent agreement is strong evidence that the structure is correct (van de Streek and Neumann, Reference van de Streek and Neumann2014). This discussion concentrates on the DFT-optimized structure. The asymmetric unit (with atom numbering) is illustrated in Figure 5. The best view of the crystal structure is down the b-axis (Figure 6). The crystal structure consists of corrugated layers of molecules parallel to the bc-plane.

Figure 3. Comparison of the synchrotron pattern of vismodegib (black) to that reported by Parthasaradhi Reddy et al. (Reference Reddy, K, Reddy, Muralidhara Reddy, Srinivas Reddy and Vamsi Krishna2014; green) for the prior art Form I, which we identified as the stable Form B. The patent pattern, measured using Cu Kα radiation, was digitized using UN-SCAN-IT (Silk Scientific, 2013), and converted to the synchrotron wavelength of 0.458968 Å using JADE Pro (MDI, 2022). Image generated using JADE Pro (MDI, 2022).

Figure 4. Comparison of the Rietveld-refined (red) and VASP-optimized (blue) structures of vismodegib. The rms Cartesian displacement is 0.055 Å. Image generated using Mercury (Macrae et al., Reference Macrae, Sovago, Cottrell, Galek, McCabe, Pidcock, Platings, Shields, Stevens, Towler and Wood2020).

Figure 5. The asymmetric unit of vismodegib, with the atom numbering. The atoms are represented by 50% probability spheroids/ellipsoids. Image generated using Mercury (Macrae et al., Reference Macrae, Sovago, Cottrell, Galek, McCabe, Pidcock, Platings, Shields, Stevens, Towler and Wood2020).

Figure 6. The crystal structure of vismodegib, viewed down the b-axis. Image generated using Diamond (Crystal Impact, Reference Putz and Brandenburg2022).
All of the bond distances and bond angles fall within the normal ranges indicated by a Mercury/Mogul Geometry check (Macrae et al., Reference Macrae, Sovago, Cottrell, Galek, McCabe, Pidcock, Platings, Shields, Stevens, Towler and Wood2020). The torsion angles involving rotation about the C20–C22 bond are flagged as unusual. These represent the orientation of the pyridine ring with respect to one of the chlorinated phenyl rings, and indicate that the overall conformation of the molecule in the solid state is unusual.
Quantum chemical geometry optimization of the vismodegib molecule (DFT/B3LYP/6-31G*/water) using Spartan ‘18 (Wavefunction, 2020) indicated that the observed conformation is 5.8 kcal mol−1 higher in energy than the local minimum, which has more-normal torsion angles about the C20–C22 bond and a different orientation of the methylsulfone group. A conformational analysis (MMFF force field) indicates that the minimum-energy conformation is 1.9 kcal mol−1 lower in energy, with further differences in the C20–C22 rotation and the different orientation of the methylsulfone group. Intermolecular interactions seem to be important in determining the solid-state conformation.
Analysis of the contributions to the total crystal energy of the structure using the Forcite module of Materials Studio (Dassault, 2021) suggests that the intramolecular deformation energy is dominated by angle distortion terms. The intermolecular energy is dominated by electrostatic attractions, which in this force field analysis include hydrogen bonds. The hydrogen bonds are better analyzed using the results of the DFT calculation.
There is only one classical hydrogen bond in the structure (Table I), N7–H31⋯N8. This is between the amide nitrogen atom and the N atom of the pyridine ring. Pairs of these hydrogen bonds link the molecules into dimers, with a graph set (Etter, Reference Etter1990; Bernstein et al., Reference Bernstein, Davis, Shimoni and Chang1995; Shields et al., Reference Shields, Raithby, Allen and Motherwell2000) R2,2(14)>a > a (Figure 7). Five C–H⋯O and two C–H⋯Cl hydrogen bonds also contribute to the lattice energy.

Figure 7. The hydrogen-bonded dimers of vismodegib, linked by two strong N–H⋯N hydrogen bonds. Image generated using Mercury (Macrae et al., Reference Macrae, Sovago, Cottrell, Galek, McCabe, Pidcock, Platings, Shields, Stevens, Towler and Wood2020).
TABLE I. Hydrogen bonds (CRYSTAL17) in vismodegib.

The volume enclosed by the Hirshfeld surface of vismodegib (Figure 8; Hirshfeld, Reference Hirshfeld1977; Turner et al., Reference Turner, McKinnon, Wolff, Grimwood, Spackman, Jayatilaka and Spackman2017) is 478.87 Å3, 96.31% of 1/4 the unit cell volume. The packing density is thus fairly dense. The only significant-close contacts (red in Figure 8) involve the hydrogen bonds. The volume/non-hydrogen atom is larger than normal at 18.4 Å3, reflecting the presence of the two Cl atoms.

Figure 8. The Hirshfeld surface of vismodegib. Intermolecular contacts longer than the sums of the van der Waals radii are colored blue, and contacts shorter than the sums of the radii are colored red. Contacts equal to the sums of radii are white. Image generated using CrystalExplorer (Turner et al., Reference Turner, McKinnon, Wolff, Grimwood, Spackman, Jayatilaka and Spackman2017).
The Bravais–Friedel–Donnay–Harker (Bravais, Reference Bravais1866; Friedel, Reference Friedel1907; Donnay and Harker, Reference Donnay and Harker1937) morphology suggests that we might expect blocky morphology for vismodegib, with {001} as principal faces. A second-order spherical harmonic preferred orientation model was included in the refinement. The texture index was 1.018(0), indicating that preferred orientation was not significant for this rotated capillary specimen. The powder pattern of vismodegib from this synchrotron data set has been submitted to ICDD for inclusion in the Powder Diffraction File.
IV. DEPOSITED DATA
The Crystallographic Information Framework (CIF) files containing the results of the Rietveld refinement (including the raw data) and the DFT geometry optimization were deposited with the ICDD. The data can be requested at pdj@icdd.com.
ACKNOWLEDGEMENTS
The use of the Advanced Photon Source at Argonne National Laboratory was supported by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357. This work was partially supported by the International Centre for Diffraction Data. We thank Lynn Ribaud and Saul Lapidus for their assistance in the data collection.
CONFLICT OF INTEREST
The authors have no conflicts of interest to declare.












