What is Evo-Devo?
The two great creative processes of biology are evolution and development. You and I, as adult human beings, are products of both. Evolution took about four billion years to make the first human from a unicellular organism that emerged from the primordial soup. Development, in the form of embryogenesis together with its post-embryonic counterpart, takes less than 20 years to produce an adult human from a different unicellular organism – a fertilized egg or zygote. By this measure, development operates more than 200 million times faster than evolution. However, despite their very different timescales, the two great creative processes of biology are intrinsically interwoven. Evo-devo is the scientific study of this interweaving. Its full name is evolutionary developmental biology, but because this is an unwieldy phrase it is almost universally referred to by its nickname.
Fundamental to any field of science is the search for general, rather than piecemeal, explanations. However, we can only generalize as far as is consistent with the facts at hand. Biology is less fertile scientific ground for generalizations than physics, because there are nearly always exceptions to any proposed general rule (for example, there are exceptions to Mendel’s ‘laws’ of inheritance). The solution to this problem for biologists is not – of course – to abandon the quest for general explanations, but rather to recognize how far any proposed generalization can go, and where its limits are set.
Against this background, we should consider the scope of evo-devo, and of any proposed general explanations that emerge from this relatively new scientific discipline. I said in the opening paragraph that evolution and development are ‘intrinsically interwoven’. However, while this is true for parts of the living world, it is not true of others. If we restrict ‘development’ to multicellular organisms in which changes occur in populations of self-adhering cells, such as embryos, larvae, or juveniles, then life forms that are unicellular throughout their entire life cycle do not have development as such. For example, a bacterium that lives for a certain period as a single metabolizing cell and then splits into two identical daughter cells by asexual reproduction cannot be said to have a developmental phase in its life cycle. In contrast, a human, a dolphin and a butterfly most certainly do have a developmental phase – indeed the butterfly has three of them (embryogenesis, larval growth, and metamorphosis).
Although I have contrasted bacteria with animals to make this point, the difference between the realms of life that are characterized by (a) occurrence of development and (b) absence of development is more complex than this introduction to the subject suggests. The realm to which development (and hence also the evolution of development) applies is that of multicellular organisms – or, to put it more precisely, the realm of organisms that go through at least one multicellular phase in their life cycles. This means that evo-devo concerns itself not just with animals but also with plants, and with some members of other kingdoms – for example the fungal and brown-algal kingdoms. It also potentially deals with certain ‘microbes’ (an undefined but generally useful term) – the ones that have a phase in their life cycle that is multicellular, albeit relatively simple, such as those cyanobacteria (previously called blue-green algae) that can form filaments or mats of attached cells.
At this point it becomes helpful to have some idea of the broad-scale structure of the living world, in terms of its hierarchical division into its three major groups (called domains) and the major subgroups within these (called kingdoms). Our view of broad-scale structure has changed considerably over the last few centuries. It will continue to change in the future, but probably much less than in the past, assuming that we are gradually homing in on a correct understanding of the course that evolution has taken. Figure 1.1 shows the broad structure of the living world, as currently perceived. Development and evolution of development characterize one of the eukaryote kingdoms in its entirety (animals), most of another (plants, defined to include both green algae and land plants), and parts of at least two others (fungi, which includes unicellular yeasts as well as multicellular toadstools, and what I think of as ‘the kelp kingdom’, which includes unicellular diatoms as well as multicellular brown algae). Other eukaryote kingdoms, and the domains of Bacteria and Archaea, are not entirely without development, but its occurrence is very patchy, and evo-devo to date has largely omitted consideration of members of these groups.
Evo-devo began in the early 1980s with studies on animals, and consequently the evo-devo of animals is better known than that of other relevant kingdoms, with the evo-devo of plants coming second. Partly because of this, and partly because I am a zoologist and know the animal kingdom better than I know the others, animal examples will predominate in this book. I hope the reader will forgive this bias, and in mitigation I can at least say that the case studies discussed will range widely across both vertebrates and invertebrates.
Although evo-devo is inapplicable to some life forms on Earth, it may well in the future turn out to be applicable to many life forms elsewhere. At the outset of his 1992 book entitled Natural Selection, the American biologist George C. Williams states a philosophical position: he believes that natural selection will be seen to characterize all life forms in however many biospheres exist in the universe – probably a very large number. Similarly, I will state a philosophical position here: evo-devo will be seen to be relevant to all life forms everywhere that are multicellular in their construction. This is a slightly different sort of statement, since natural selection is a process while evo-devo is a field of study. However, I would predict that the most important processes involved in the evolution of development here, as discussed in later chapters, will be relevant on other planets too.
We’ve now considered the realm within which evo-devo studies are meaningful. Or, in other words, we’ve clarified the realm in which evolution and development are interwoven. Having done that, we now turn to the way in which they are interwoven. From here on I will focus on the animal kingdom, unless specified otherwise. However, many of the points made will be equally applicable to plants, and to other multicellular organisms.
The development of any animal can be thought of as a trajectory from zygote to adult. Since I will be using ‘developmental trajectory’ often in this book, I should explain here what I’m thinking about when I use this term. Imagine the development of a human. Each of us starts our life as a single cell, which starts to multiply, producing a self-adhering cluster or ball of cells. As it continues to grow, this cluster begins to take a more definite shape, with elongation in one direction producing the anteroposterior (or head-to-tail) axis of the body. Internal changes, such as the separating out of different tissue layers, accompany the external changes in shape, and the overall growth. The embryo gradually elaborates its features, becoming more and more like a miniature human. After birth, development continues, but is more subtle. One important aspect of the post-embryonic development of a human is differential growth rates of different parts of the body, something that is referred to as allometric growth (to distinguish it from isometric growth, where different body parts grow at the same rate). For example, our heads grow more slowly, after birth, than our trunks and limbs. The combination of all these changes leading from zygote to adult constitutes the developmental trajectory of a human.
At any moment in evolutionary time – say halfway through the Jurassic period – the developmental trajectory found in a certain kind of animal – say a particular species of dinosaur – has been produced by the accumulated evolution of the past and is the starting point for the evolution of the lineage concerned in the future. Development is a quasi-predictable process. Its many repeat occurrences within a given species produce variants that are typically rather similar to each other – though not identical. In contrast to development, evolution is a very unpredictable process. The fact that one dinosaur developmental trajectory gave rise to all of today’s 10,000 species of birds while the others left no descendants among today’s fauna could not have been foretold. Evolution incorporates a major element of ‘historical contingency’ – chance events including asteroid impacts and volcanic eruptions – as emphasized by the American palaeontologist Stephen Jay Gould. Development is usually much less affected by such contingency.
One way of looking at the intertwining of evolution and development, then, is that development is a sort of raw material that gets moulded by natural selection in ways that adapt it to the prevailing environmental conditions in the relevant habitat. For example, most frogs, including all those living in temperate habitats, have a process of indirect development that includes a tadpole stage, but some species living in warm, moist, tropical conditions have dispensed with the tadpole stage of this ancestral life cycle and have evolved a process of ‘direct development’ (the embryo gives rise directly to a juvenile that’s like a small adult). Among the many species of frogs living in temperate regions, evolution has again moulded the pattern of development to fit the environment, but in less dramatic ways. For example, a shorter tadpole phase of the life cycle would be expected to be favoured by selection in regions where the water bodies inhabited by the tadpoles are more transient than they are in others.
But the interweaving of evolution and development is not a one-way street. Evolution alters the developmental process, to be sure. But the evolutionary process is also altered by development. Or, to put it another way, evolution is effectively channelled in terms of what it can do with a particular lineage in the future by the prevailing developmental trajectory of the species concerned in the present. Likewise, evolution at any point in the past – say the mid-Jurassic again – was channelled by the developmental trajectories that were available at that time. This channelling is often referred to as ‘developmental constraint’. However, I think this phrase has an overly negative tone. If development in some sense channels the direction of evolution, then it can be thought to steer it towards some types of change just as much as it steers it away from others. We’ll look at various examples of this steering effect, which can be called developmental bias, in subsequent chapters.
The practitioners of evo-devo are not a homogeneous bunch. Those who are above a certain age have migrated there from various disciplinary backgrounds, because when they were students the field did not exist – or perhaps had just begun but was not yet widely studied or well funded (regrettably the last of these remains true, though the situation is a little better than it used to be). Some practitioners have come from developmental biology, some from evolutionary biology, and some from elsewhere. Partly because of their heterogeneous origins, these practitioners are also rather heterogeneous in their views of the nature of evo-devo. With regard to the two sides of the interaction between evolution and development noted above, some emphasize one, some the other, and some both. This heterogeneity of views is of interest in relation to the wider philosophy of evo-devo. However, before considering the philosophical aspects of the new discipline, we need to know a little more about it – and that includes understanding its origins.
Origins of Evo-Devo: the Homeobox
If you were to ask me, ‘what was the single most important discovery in the origin of evo-devo?’ I’d reply, with little hesitation, ‘the homeobox.’ So that’s a good place to start. This was a discovery made at the same time (in 1983, with publications following in 1984) by researchers in two laboratories – that of Thomas Kaufman in Bloomington, Indiana, and that of Walter Gehring in Basel, Switzerland. The lead authors of the papers concerned were Matthew Scott and William McGinnis.
The key question at this point is: what is a homeobox? To answer this we need to start with the related question: what is a gene? A reasonable working definition is that a gene is a stretch of DNA that’s thousands of nitrogenous bases long, and that makes a particular product (typically a protein). Each different gene makes a different protein, because each gene is a unique sequence of the four bases that we’re familiar with by their initial letters of A, C, G, and T (full names adenine, cytosine, guanine, and thymine). Recall that the genetic code works in triplets, so three bases in a DNA strand give rise to one amino acid in the resultant protein. For example, the sequence AAA in a gene corresponds to the amino acid lysine in the protein. If we say that a typical protein is 333 amino acids long (just a rough guess), then the gene making it must be at least 999 bases long. In fact, it is normally much longer than this for various reasons, principally that the genes of organisms from the kingdoms that have development (animals, plants, etc.) typically contain stretches of DNA (called introns) whose RNA counterparts are chopped out before the protein is made. So the typical animal or plant gene is in fact thousands of bases long, rather than hundreds.
Now we return to the homeobox. A ‘box’, in genetics, is simply a rectangle drawn around a certain stretch of DNA to highlight it, for whatever reason. For example, if I wanted to highlight the AAA stretch in the following sequence to show you which bit of a longer sequence coded for the amino acid lysine, I’d draw a box around the central three bases (here I’m using bold text to do the same thing): TATAAAGGG. The homeobox is a much longer sequence of bases than this. It is typically 180 bases in length, thus corresponding to a sequence of 60 amino acids in the corresponding protein – which in turn is called the homeodomain. In a homeobox-containing gene whose overall coding sequence is 1800 bases long, the homeobox is 10% of the gene (Figure 1.2), whereas in a gene that is 18,000 bases long (perhaps due to multiple long introns), the homeobox is just 1% of the gene’s complete DNA sequence. In a homeodomain-containing protein that is 300 amino acids long, the homeodomain itself makes up 20% of this overall length.
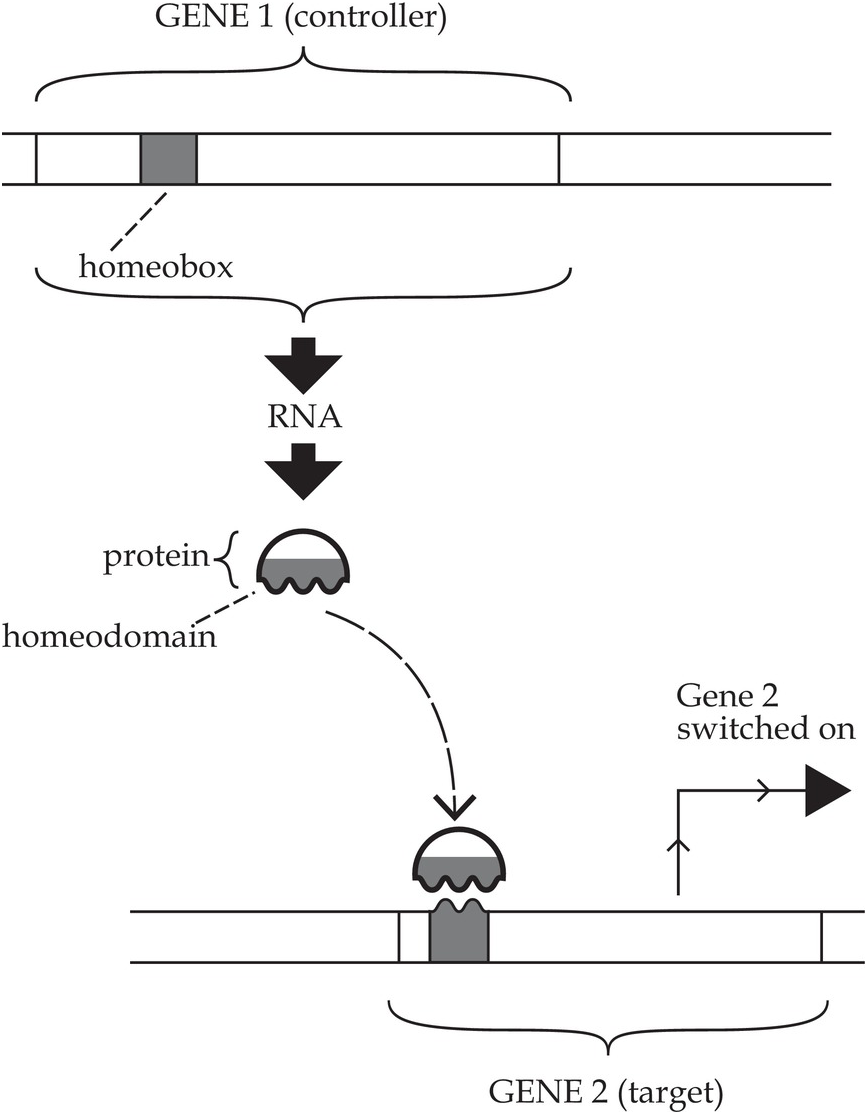
Figure 1.2 Diagram of a homeobox gene and the homeodomain protein that it makes. The homeobox represents only a small fraction of the overall length of the gene. Likewise, the homeodomain represents only a fraction of the protein. The important point to note here is that the protein’s homeodomain enables it to bind to the DNA of other genes and to switch them on, thus causing them to make their own protein product. This provides a basis for developmental patterns of gene activity in the embryo.
So far, we recognize a homeobox as a sequence of a particular length that can be found within certain genes. But what is the sequence, which genes is it found in, and why is its inclusion in these genes significant?
It’s not possible to specify the 180-base homeobox sequence exactly, because there are many variant versions of it. It’s a recognizable pattern, or ‘motif’, rather than a precise sequence. Typically, one variant will be the same as another for most of the 180 bases, but will differ in a minority of them. Part of this variation is due to the redundancy of the genetic code. For example, it’s not just AAA that codes for the amino acid lysine, AAG does so too. Thus it’s possible to get a homeodomain with lysine in a particular position, by having either of these triplets in the corresponding stretch of the homeobox that codes for it.
But there is variation in the exact amino-acid sequence of homeodomains too. One variant homeodomain will typically be the same as another for most but not all of its amino acid sequence. The most important thing is that regardless of some variation in the structure of the homeodomain at this level, at a higher level its overall 3D structure is maintained. This 3D structure is much more complex than shown in Figure 1.2, which is schematic. We don’t need to know this structure in detail, but its key feature is that it has three helical regions that are vital to its DNA-binding function, and that are conserved in all variants.
Now let’s turn to the genes in which a homeobox sequence is found. Genes can be roughly divided, in terms of their function, into three main categories: developmental (crucially important here), cell-type specific (less important), and housekeeping (least important in evo-devo). As the names of these rough-and-ready categories imply, the first make proteins that contribute to the developmental process, the second make proteins that are restricted to certain cell types (such as the haemoglobin of our red blood cells), and the third make other proteins (often enzymes) that contribute to the housekeeping tasks that occur within almost all cells, such as the metabolic activities that keep a cell supplied with energy. As usual in biology, categories are never clear-cut, but this way of thinking about gene function is helpful nevertheless – and we’ll delve further into it in Chapter 4.
Many developmental genes contain a homeobox; in contrast, other genes generally lack this sequence. There is a good reason for this difference. A major part of the causality of development is cascades of gene activity in which the product of gene A switches on gene B, whose product switches on gene C, and so on. To switch on a gene, a protein must bind to the DNA of that gene. And it turns out that the homeodomain is a DNA-binding region. So if we discover a new gene about which we initially know nothing, sequence it, and discover that it contains a homeobox, we are pretty sure that it plays a role in the development of the animal concerned.
The animal in which the homeobox was discovered was the fruit-fly Drosophila melanogaster. The genes in which it was discovered were already known from the fact that mutations of them produce bizarre adult flies that have legs growing out of their heads or two pairs of wings instead of the normal single pair. Back in the late nineteenth century, these mutations were called homeotic, and the phenomenon they produce was called homeosis – the right thing in the wrong place. This contrasts with the wrong thing in the right place, which is a much commoner type of mutation. An example of the latter in Drosophila is the vestigial-winged mutant, where the wings are in the right place but are small and malformed. The person who coined this terminology (homeosis/homeotic) was the British geneticist William Bateson, whose book Materials for the Study of Variation was published in 1894. We’ll come back to him in Chapter 2, when we look at the antecedents of evo-devo. For now, it’s just important to know that it was from his homeosis that the homeobox sequence got its name.
The huge significance of the homeobox arises from the fact that it was found to characterize multiple developmental genes and, as research continued in the 1980s, it was found to characterize such genes not just in flies but in animals generally. This suggested that the genes that contributed to the developmental process were similar even when the end result of the process – the adult animal – was very different. In other words, it suggested that we could generalize about some aspects of the causality of development right across the animal kingdom, and perhaps even beyond. To say that this was exciting stuff would be an understatement. However, one discovery does not by itself make a new scientific discipline. So let’s now look at some important events that preceded and followed the discovery of the homeobox.
Origins of Evo-Devo: Other Factors
In 1977, Stephen Jay Gould rekindled interest in the relationship between evolution and development with the publication of his book Ontogeny and Phylogeny (which, with a few ifs and buts, simply means Development and Evolution). Two years later, he and fellow Harvard biologist Richard Lewontin wrote an influential – and controversial – paper championing the role of various forms of constraint in evolution, including developmental constraint, and downplaying the role of natural selection. We’ll discuss this paper, and the concept of constraint, in Chapter 4.
In 1978, the American geneticist Ed Lewis published an important paper on the structure of a gene complex in Drosophila that contained multiple genes that were subject to homeotic mutation and that would, a few years later, be found to contain homeoboxes. In 1980, the Heidelberg-based biologists Christiane Nüsslein-Volhard and Eric Wieschaus published an equally important paper on other genes that contribute to the development of Drosophila, including the now-famous hedgehog gene that gives mutant larvae a prickly appearance – hence the name. The genes studied by these three biologists were similar in that they all affected the development of segments – those sections of the body along the head-to-tail axis of which an insect is made. However, in another way they were different. The genes studied by Lewis were involved in the determination of segment identity (e.g. thoracic vs. abdominal). In contrast, those studied by Nüsslein-Volhard and Wieschaus were involved in the determination of segment number and segment polarity (which end of a segment is anterior and which posterior); these authors showed that such determination took place in stages of increasingly restricted spatial scope, starting with the whole body and culminating in the patterning of individual segments. All three of these biologists shared a Nobel Prize in 1995 for their work on developmental genes, which helped to pave the way for later comparative studies.
Through the rest of the 1980s, the 1990s, and into the present century, work on comparative developmental genetics expanded rapidly. Other developmental genes than those subject to homeotic mutations were discovered to contain homeoboxes, so the relationship between homeobox genes and homeotic genes became more complex than the one-to-one relationship that might at first have been thought. And many developmental genes were discovered not to have homeoboxes, which suggested that some of them might have a different developmental role than making proteins that directly switch other genes on. For example, the hedgehog gene makes a protein (called Hedgehog) that is secreted out of a cell, transported in the intercellular matrix, and bound by a receptor protein on the surface of another cell. Hence it is a cell–cell signalling protein – one of a number of such proteins that make up what is called a signalling pathway. This is important, because development involves not just the turning on of certain genes in certain cells but also the coordination of such switching events between cells in space and time, both in embryos and in post-embryonic developmental stages.
Although certain genes are crucially important in the developmental process, I have deliberately avoided using the statement ‘genes control development’, which regrettably can be found in many publications. Such statements are rightly criticized as smacking of what is sometimes called genetic imperialism. In fact, development is a two-way interaction between genes and other molecular players, including proteins. Genes make proteins, while certain proteins control the activities of genes; and this interplay is often influenced by environmental factors. In a collaborative study by members of Michael Akam’s laboratory in Cambridge and my laboratory in Galway, it was shown that in contrast to the situation in flies, where the number of segments is not influenced by the environment, the number of segments in some species of centipedes can be altered by the temperature at which embryos develop – this finding was published by Vincent Vedel and colleagues in 2008. A year later, the American biologists Scott Gilbert and David Epel published their book Ecological Developmental Biology (second edition 2015), which emphasized the importance of the environment – including other species – in influencing the route that development takes. The nickname eco-devo is now sometimes used for such an approach.
Evo-Devo and Darwinism
In The Origin of Species, first published in 1859, Darwin famously described natural selection as ‘the main but not exclusive means of modification’ in the evolutionary process. While some version of this statement is probably true, it’s not as simple a statement as it first seems. To see why, we need to discuss levels of biological organization. These extend from the molecular level up to that of the cell, then to tissues and organs, then to the individual animal or plant, and finally (for our purposes here) up to the population – the collection of all those individuals of a particular species living in a particular area.
Natural selection is a population-level process. At this level, selection probably is, as Darwin suggested, the main means of evolutionary change. Perhaps the only evolutionary theory of the last half-century or so that denies this is the neutral theory, proposed by the Japanese geneticist Motoo Kimura in the 1960s and explained at length in his 1983 book The Neutral Theory of Molecular Evolution. Kimura argues that, with certain provisos, more of the evolutionary changes in genes and their protein products are driven by ‘genetic drift’ than by natural selection. Drift is the name given to the random fluctuations in a population’s genetic make-up that are most noticeable when selection is absent. In the short term, when applied to a single gene, these don’t achieve much; however, in the long term, across many genes, drift will sometimes cause many changes in the same direction, just by accident. It’s like asking all the students in a lecture room to toss a coin ten times. If there are enough students, someone will get ten heads in a row just by chance.
However, the ‘certain provisos’ mentioned above are important. While Kimura questioned Darwin’s theory at the level of individual genes, he acknowledged that, in terms of organismal body form, natural selection prevails. So, the long neck of a giraffe is not a result of genetic drift any more than it is a result of the giraffes’ neck-stretching being passed on to their offspring (the so-called ‘inheritance of acquired characters’, for which there is no evidence).
Evolution is a multi-level process, not just a population process. It involves molecules, cells, and organisms, as well as populations. What happens when we simultaneously consider more than one level? Here are two alternative multi-level scenarios.
1. Suppose that many mutationally induced changes to development are possible, and that they are all equiprobable. If a new variant is fitter in a general sense, as summed up over all stages of the life cycle, it may spread under the influence of natural selection in the population. In such a scenario, there is no conflict between developmental and population levels of organization.
2. Suppose instead that some variant patterns of development are inherently more probable than others in the sense of being more easily produced (this can be called developmental bias, arguably a better term for developmental constraint, as noted earlier). Bias could arise because of some aspects of the dynamics of the developmental process. If it is strong enough, bias in favour of one variant may be more important than natural selection in favour of the alternative variant.
The Canadian biologist Brian Goodwin suggested the second scenario in the case of leaf-development patterns in flowering plants (the angiosperms: a group of more than a quarter of a million species). There are three main such patterns: (a) leaves branching off a stem on alternate sides (distichous); (b) leaves branching off in pairs or groups, with each group being rotated through 90 degrees with respect to the previous one (decussate); and (c) each leaf branching off at a certain angle to the one before it (spiral). In his 1992 book How the Leopard Changed its Spots, Goodwin notes that more than 80% of angiosperm species have the spiral pattern of leaf development. And he goes on to suggest that this is because the spiral pattern is the most easily generated by plant developmental systems. He says (Chapter 5, page 132): ‘the frequency of the different phyllotactic patterns in nature may simply reflect the relative probabilities of the morphogenetic trajectories of the various forms and have little to do with natural selection.’
There are, however, at least two problems with Goodwin’s suggestion. First, the significance of 80% of angiosperms having a spiral pattern of leaf development depends on how they are scattered across the angiosperm evolutionary tree. The spiral developers could have had just one origin, and characterize a single but very large branch. Alternatively, they could have arisen lots of times and characterize instead multiple small twigs. Goodwin did not deal with this aspect of the problem, and perhaps in retrospect he was wise – because the perceived structure of the angiosperm phylogenetic tree changed dramatically with the blossoming of plant molecular phylogenetics shortly after his book was published.
The second problem is this: it’s hard to imagine that the pattern of arrangement of the leaves of an angiosperm would not be subject to natural selection. After all, leaves are the major energy-acquisition organs of flowering plants (with some exceptions, such as cacti). The pattern in which they develop and hence are arranged thereafter must be of major importance in terms of minimizing leaf overlap and maximizing photosynthetic activity. However, Goodwin suggests (on page 132 again) that ‘all the phyllotactic patterns may serve well enough for light-gathering by leaves and so are selectively neutral’. It is one thing to suggest, as Kimura did, that a single amino-acid change in a protein that contains hundreds of amino acids may be selectively neutral; it is quite another thing to suggest that the overall structure of a plant’s main photosynthetic apparatus is neutral.
Clearly, the philosophical stances associated with advocating the two scenarios described above are very different. Number 1 is essentially ‘neo-Darwinian’ while number 2 could be described as ‘anti-neo-Darwinian’. But there is an intermediate scenario, with an associated intermediate philosophical stance, which will gradually come into focus as we proceed. In this scenario, developmental bias and natural selection interact with each other to produce evolutionary trends in particular directions. Many practitioners of evo-devo are attracted by this idea. (An interesting source of information on the role of developmental bias in evo-devo thinking is provided by a 2020 special issue of the journal Evolution & Development, edited by Armin Moczek.)
However, many practitioners of evo-devo are not overtly philosophical in their approach. I would describe the molecular evo-devo that began in the 1980s as a largely practical endeavour, in which the focus was on doing more case studies, accumulating more information in general, and waiting to see what generalizations suggest themselves when we have a more extensive database. Today, much evo-devo is still of this data-gathering flavour. It is particularly important in this endeavour that our data are well spread across the animal and plant evolutionary trees. This can be referred to as good taxon sampling, both in the sense that no major branches are omitted and in the sense that we can make multiple comparisons at all levels of taxonomic distance, from closely related species (e.g. those belonging to the same genus or family) to representatives of different phyla.
Despite its current practical focus, evo-devo is not a philosophy-free zone. Indeed it has attracted the attention of many philosophers of science, who are interested in the impact that evo-devo may have on our understanding of evolution in general. Of course, it might turn out that many evo-devo studies are just filling in some missing details. Perhaps this is true of those studies that revealed some of the developmental genes and proteins that underlie the differences in beak size and shape among different species of Darwin’s finches, without altering the basic adaptive scenario under which the beaks are thought to have evolved. Then again, it may turn out that some evo-devo studies are really saying something new and important about how evolution works. This is perhaps most likely to be the case in relation to the origins of evolutionary novelties and body plans (e.g. the turtle shell and the vertebrate skeleton respectively). We will deal with these origins from various perspectives in Chapters 6, 7, and 8.
The Importance of Evo-Devo
Evo-devo has a general importance that transcends the issue of whether it does or does not end up producing a new view of the ways in which evolutionary novelties and body plans originate. Our new discipline is essentially putting the individual animal or plant back at centre-stage in our overall perspective on the pattern and process of evolution.
In the couple of decades before evo-devo started, much of evolutionary theory was focused on two particular levels of organization – molecules and populations. Strangely, the intermediate level of the individual organism was neglected. The prevailing approach in the field of population genetics was that we could be dealing with a barnacle or an elephant – it didn’t really matter. Although this approach now seems bizarre, it arose for a good reason. It’s important to understand this reason, because it helps us to put the evo-devo endeavour that came later (the 1980s) into a broader evolutionary context. And to understand why molecular population genetics became so dominant in the 1960s and 1970s, we need to look back a couple of decades earlier than that.
In the 1940s and 1950s, evolutionary case studies carried out in the wild were typically based on phenotypic variation in natural populations that was visible to the naked eye. This was because many of the molecular methods that are now used to ‘see’ things – like genes and proteins – inside organisms did not yet exist. Some studies focused on continuous variation – such as that in the size of a snail’s shell. However, the interpretation of such studies was complicated by the fact that continuous variation is only partially inheritable. Other studies focused on polymorphic variation, that is, the type of variation in which there is a small number (sometimes just two) of phenotypes or ‘morphs’ that are clearly distinct from each other and do not intergrade via a long series of intermediates. An example is the case study of different colours and banding patterns on the shells of various species of land-snails, notably Cepaea nemoralis, and the way in which the relative frequency of these morphs varies from place to place. This latter type of study had the advantage that breeding experiments revealed the discrete phenotypes to be completely inheritable and controlled by particular genes. However, they had the disadvantage that most animals do not exhibit polymorphic variation that is externally visible. Hence the case studies were restricted to groups that do – land-snails, lepidopterans, ladybirds, and a few others. Their broader relevance was uncertain.
In the 1960s everything changed. The introduction of a particular technique (gel electrophoresis) allowed the screening of natural populations of any type of organism for genetic variation in the many genes that produce enzymes. Pioneering work on populations of flies and humans revealed that such genes were often polymorphic. Since all organisms have multiple enzyme-producing genes, this work could be (and was) extended to many taxa. Unlike the earlier work on pigmentation polymorphisms, the work on enzyme polymorphisms was clearly applicable across the living world. And the main discovery was that polymorphism was widespread. By the 1970s, this had been established beyond reasonable doubt.
This discovery was followed by a heated debate over whether variation within and between populations was acted upon by natural selection, or was selectively neutral and hence only acted on by the random process of genetic drift. The answer turns out to be ‘a bit of both’, with the relative importance of the two still not entirely clear. Two of the key books about this debate are Richard Lewontin’s The Genetic Basis of Evolutionary Change, published in 1974, and Motoo Kimura’s 1983 book The Neutral Theory of Molecular Evolution, which I mentioned earlier.
With the benefit of hindsight, this work of the 1960s and 1970s can be thought of collectively as being ‘the evolutionary genetics of housekeeping genes’. Many of the enzymes studied were standard players in metabolism, for example enzymes involved in the energy-generating processes that are common to most cell types, such as glycolysis (liberation of energy from glucose). Hence the loss of focus on the organism and its body form. The developmental trajectories of a barnacle and an elephant are very different, but their main metabolic pathways are much the same.
In the 1980s, interest shifted in two directions. Studies of molecular evolution shifted from gene products – proteins – to the genes themselves, and these studies branched out to incorporate evolutionary change in DNA sequences that are not ‘standard genes’ – e.g. repetitive DNA, transposable elements, and pseudogenes. (To learn more about that work, you will need to read an introductory book on molecular evolution, as it’s outside the scope of this one.) At the same time, evo-devo arose from discoveries such as the homeobox, which allowed us to examine not housekeeping genes but developmental genes – the genes that build the organism.
So, the importance of evo-devo, at its most general, is that it has put the organism back at the centre of evolutionary thinking. We are not just interested in the evolution of an animal’s metabolism, we are also interested in the evolution of its form. And indeed perhaps we should be more interested in the latter, though of course this is a subjective opinion. It is an opinion that was nicely put by the British biologist Conrad Hal Waddington (whom we’ll meet again in Chapter 2). In 1975 – shortly before the advent of evo-devo – Waddington perspicaciously said this:
It is doubtful if anyone would ever have felt any need to resist the notion of evolution if all it implied was that the exact chemical composition of haemoglobin changed over the ages.
Groups of people resisting the idea of evolution – biblical fundamentalists and other creationists – do so on several bases. One is the unacceptability (to them) of evolutionary theory’s refutation of the creation myth featuring Adam and Eve, some version of which appears in all the Abrahamic religions – Judaism, Christianity, Islam, and several other smaller ones. Another is the notion of the supposedly ‘irreducible complexity’ of biological systems – the idea that these systems are too complex to have arisen by the blind processes of mutation and selection. While molecular complexity (e.g. Waddington’s haemoglobin) is in fact objected to as a product of evolution by creationists, what they object to much more is the idea that objects as impressively complex as the teeth of a tyrannosaur or (especially) the brain of a human are also products of evolution.
We can link this argument about what creationists object to the most with the question of what would excite biologists the most in terms of the discovery of life on other planets. Finding a biosphere consisting entirely of microbes would be very exciting – and such a discovery may only be a few decades away (via studies of exoplanet atmospheres). But finding a biosphere with large multicellular creatures characterized by complex developmental systems that produce, among other things, sophisticated brains, would be more exciting still. It is evolution’s pinnacles of achievement that excite us most of all.
But let me finish with a caveat. Evolution is not an escalator. There is no such thing as a universal upward trend in brain size (or anything else), as we will discuss further in the next chapter, in terms of the old idea of a scala naturae. In the lineage from an ancient worm to a modern human, brain size started small and then got bigger and bigger, but this trend was interrupted by frequent long periods of stasis and perhaps also by shorter periods of decline. One of the most closely related animal phyla to the vertebrates is the Echinodermata – the group that contains the starfish, sea-urchins, and sea-cucumbers, among others. In the lineage from that same ancient worm (the last common ancestor of vertebrates and echinoderms) to a present-day starfish, brain size went from small (possibly via a bit bigger) to zero. We always need to keep the messiness of evolution at the back of our minds. Evolution isn’t ‘trying’ to achieve anything. It simply happens. It’s an inevitability of having the three linked properties of variation, reproduction, and inheritance. And the evolution of development is an inevitability of having those three plus the fourth property of having a multicellular body with a developmental trajectory.



